ANGIOSPERM PHYLOGENY WEBSITE, version 14.
On classifications in general, and in particular on the classification used here.
On ancient hybrids, introgression, etc. — and phylogenetic trees with grafted branches.
On forming clade characterizations (and thinking about apomorphies).
SUMMARY OF APG IV, MAIN CHANGES SINCE, LINKS TO PAGES.
On some poorly-known taxa that are in need of study.
On the organization and design of this site.
On the interpretation of the text, etc.
Important - Warning to All Users!
Website originally developed by Hilary Davis.
Systematics is a profoundly historical discipline, and we forget this at our peril. Only with a phylogeny can we begin to understand diversification, regularities in patterns of evolution, or simply suggest individual evolutionary changes within a clade. Our recovery of that phylogeny is the recovery of evidence of a series of unique events that comprises the history of life. These pages are a series of characterizations of all orders and families of extant angiosperms (flowering plants) and gymnosperms, i.e. all seed plants, as well as many of clades grouping families and orders and some smaller clades, especially within larger families; non-seed plants are covered more briefly. The pages are designed to help in teaching seed plant phylogeny at a time when our knowledge of the major clades of seed plants and the relationships within and between them are still somewhat in a state of flux, even if much of the broad outline seemed to be becoming clear (but c.f. A.P.G. IV 2016), one can see a daily-updated Tree of Life based on genome sequences as they appear, for example (Fang et al. 2013), a comprehensive Tree of Life is being developed (Hinchcliff et al. 2015; see below). Nevertheless, as of viii.2018, I estimated that there were still substantial questions about relationships in about twelve orders of seed plants, less serious questions about five more, and four or more internal nodes/areas on the main tree where there were important problems - that was just before the advent of data from comprehensive nuclear analyses. Perennial aggravations have been the position of Gnetales and relationships of and within the bryophytes, although there seems to be progress with these issues (e.g. Ran et al. 2018 and Wickett et al. 2014, Puttick et al. 2018 and Morris et al. 2018, respectively). From around 2018 onwards the almost routine analysis of massive amounts of data from chloroplast genomes in particular seemed to be shedding some light on such relationships, but now nuclear data at the whole genome/transcriptome level are questioning some relationships that we thought we understood (e.g. Sun et al. 2014: rosids; Rydin et al. 2017: Rubiaceae, mitochondria; O.T.P.T.I. 2019: focus on angiosperms; C. Zhang et al. 2020; Cai et al. 2020; W. J. Baker et al. 2021a, b, and see the associated (and developing) Seed Plant Tree of Life, papers in American J. Bot. 108(7). 2021, etc.). As details of phylogeny are clarified and new findings made in anatomy, morphology, etc. (but there are far fewer of these latter appearing than one would like), they can be rapidly integrated into the Angiosperm Phylogeny Group system that is followed here (see A.P.G. IV 2016), and you can find some of the main developments since then below. Indeed, there are discussions as how best to provide the means to produce trees and analyse features in the context of phylogenies using vast amounts of data in a repeatable way and on a regularly updatable basis (Eiserhart et al. 2018) - we shall see.
Although books are out-of-date before they appear, there is a useful comprehensive phylogeny-based treatment of angiosperms by D. Soltis et al. (2017, ed. 2); see also K. Bremer et al. (2004b). There are good treatments of the main European families in Sitte et al. (2002) and the 37th edition of Strasburger's Lehrbuch (Kadereit et al. 2014), and of North American - and many other - families in Judd et al. (2015, ed. 4) and Simpson (2019, ed. 3). Neotropikey (Milliken et al. 2009 onwards) - more than a simple interactive key - is a resource for students of the Neotropical flora while the most recent edition of Adolf Engler's Syllabus of Plant Families/Syllabus der Pflanzenfamilien (Frey 2015) that is coming out is based on recent findings in phylogeny, while Keller (2023) provides a comprehensive field guide to tropical families. There is a regularly updated and printable Plant Phylogeny Poster (now in more than 20 languages: Cole et al. 2021) and a different visualization of the seed plant universe at Botanical Chart. Johansson (2013) and Byng (2014) are comprehensive phylogeny-based accounts of angiosperms (the former is no longer being updated), Byng (2015) covers gymnosperms, while Lecointre and Le Guyader (2016) are going to cover everything. The beautifully-illustrated Christenhusz et al. (2017) includes accounts of all vascular plant families, and then, depending on what part of the world you are living in, more local treatments may catch one's fancy. Thus Koekemoer et al. (2023) have recently come out with Flowering Plant Families of Southern Africa - again, superbly illustrated.
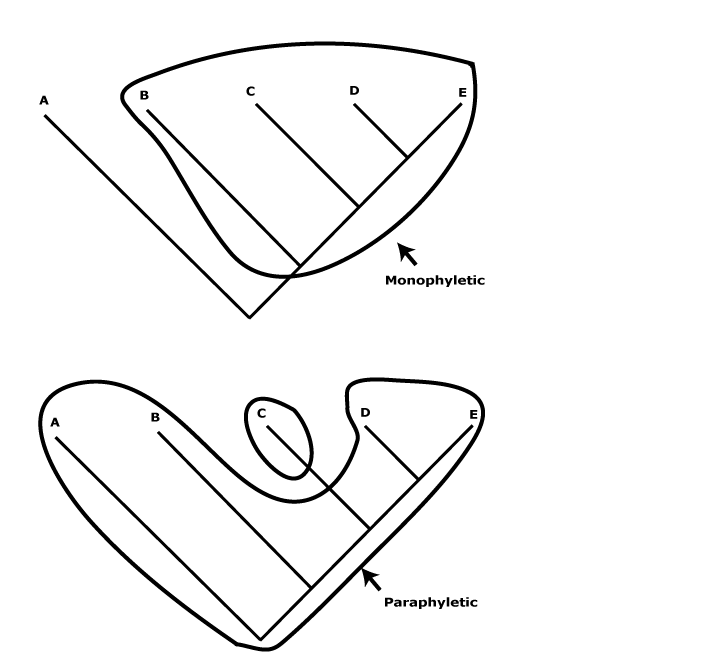
The focus of this site is on angiosperm families and in a number of instances their subfamilies/tribes, sometimes even genera, while treatments of gymnosperms were added in 2005 and other embryophytes/land plants are covered in a less detailed fashion. Emphasis is placed on plant families because they are the groups - admittedly partly arbitrary as to circumscription, but now monophyletic, i. e., including all and only the known species of a common ancestor (however, see below) - around which many of us organize our understanding of plant diversity. I also pay attention to groupings of families and orders (for which, see A.P.G. IV 2016, but see caveats above), while infrafamilial groupings in larger families like Annonaceae, Apocynaceae, Ericaceae, Fabaceae, Malvaceae, Poaceae, Rubiaceae, Rutaceae, Scrophulariaceae, etc., are being added as studies become available. I tend to emphasize literature that deals with clades with fifty or more species, although in smaller families the coverage is more detailed. Note that a World Flora Online (http://www.worldfloraonline.org) is being developed that ultimately will include accounts of all species (Borsch et al. 2020). This more or less represents a concensus over species limits (see also below) and it should be consulted for information.
Relationships between families, etc., are shown as branching diagrams or trees; Baum and Smith (2012) is a good account of how to interpret and think about such trees - although it will become clear that simple trees may not always be the best way to represent relationships. Trees are a means to an end, that is, to help us understand - in the broadest sense - pylogeny. Throughout the site, the treatment of variation is as hierarchical as I can make it, with putative apomorphies being mentioned at the appropriate place in the tree, although in the ultimate groupings both apomorphies and characters varying within these groups may be mentioned. Finding out the composition of clades is quite often easier than finding the synapomorphies for those clades, indeed, recent developments in ancestral state reconstruction make it clear how difficult the latter can be (see the discussion below), and uncertainties about the ages of clades remain pervasive. And knowing about synapomorphies and ages, difficult although this may be, is just the beginning of understanding the whys and wherefores of the evolution and diversification of seed plants, our ultimate goal. Hence the sections on various aspects of evolution that include "Divergence & Distribution", "Ecology & Physiology", "Pollination Biology & Seed Dispersal", "Plant-Animal Interactions" and "Bacterial/Fungal Associations" in these pages. And of course phylogeny is just one way of framing our understanding of land plants, but I hope it will become clear that it is a very important one that allows the incorporation of other viewpoints, too, I hope giving them interesting new perspectives.
There is a separate and more extended discussion on the evolution and diversification of angiosperms as a whole, and here in particular the emphasis is less on species numbers, more on what species/clades "do", and the major role that plants play in constructing the environment in which they (and we) live. Here thinking about the microbiome, other organisms closely associated with plants such as bacteria and endophytic and mycorrhizal fungi, is very important, since it is the plant and microbiome together that help shape the ecological relationships between plants and their environment. Indeed, knowledge of the microbiome and its interactions with the host plant changes what we think of as an individual plant - a plant is part of a holobiont, a group of more or less closely integrated and interacting organisms, and genomes of all these organisms make up the hologenome, a microcosm - maybe not so micro - that can be thought of as the holobiome (Bordenstein & Theis 2015; Gilbert & Tauber 2016; Martin et al. 2017; Tripp et al. 2017a: symbiome). Thus the ectomycorrhizal (ECM) associates of seedlings of pinyon pines (Pinus edulis) differed, but the ECM fungi and the genotype of the seedlings resembled those of the parent pines, which were either drought-tolerant or -intolerant (Gehring et al. 2017); and as those authors (ibid.: p. 11170) noted, "ECM community composition represents a heritable plant trait". Along the same lines, the effect of a caterpillar on a plant can be manipulated by parasitoids of that caterpillar, and ultimately by polydnaviruses in the parasitoids - the saliva of the infected caterpillar is less likely to elicit plant defences than that of an uninfected caterpillar (Tan et al. 2018; see also Zhu et al. 2018) - all for the benefit of the polydnavirus.
Further information about all families mentioned on this page may readily be found by using the "Families" link in the top bar or the Summary below. General information about the characters mentioned may also be found in the characters page. But before going further, it should be emphasized how little we know about most plants (e.g. Cornwell et al. 2018/2019), and this is particularly true of details of anatomy, embryology, development, etc., that are central to this site, but whose study, with the partial exception of development, is now hardly "fashionable".
ON CLASSIFICATIONS IN GENERAL, AND THIS CLASSIFICATION IN PARTICULAR
[Back to Top]
On classifications in general.
Godfray and Knapp (2004: p. 562) observed that "users want stable, informative and accessible classifications that enable easy identification" - although invoking "users" without specifying those who make up this group rather begs the question. The classification here is for all interested in comparative biology, hence the emphasis on phylogeny; there are, as we shall see, many ways of making such a classification accessible to all.
Classifications in the broad sense are box-in-box, group-in-group, or part/whole naming devices that we use to communicate aspects of our knowledge of things in general (see Parsons & Wand 2008 for an introduction). For any classification system to be effective, it must be stable, universal, i.e., be used by a wide range of people, and it must enhance communication of knowledge by helping us to relate things in our minds. From this point of view biological classification systems are no different from any others (Stevens 2006a for references). The phylogeny-based classification used here conveys aspects of our knowledge about the phylogenetic relationships about land plants. Thus a family is clearly flagged as such and is a monophyletic group that can contain one to many genera, also flagged as such and also for the most part monophyletic, but a genus can never include families. Generic, family, etc., names are simply words we use to denote appropriate parts of phylogenies and convey some, if rather minimal, information about their relationships. The circumscription of individual taxa may as much reflect the fact that the birthplace of contemporary systematics was Europe and North America, not the Antipodes (e.g. Walters 1961), rather than any philosophical ideas of the authors describing these taxa (some nineteenth century botanists were aware of this local, eurocentric bias).
Here I use what may be called a flagged hierarchy (Stevens 2006a) for naming taxa. The only thing that the rank terminations used (-ales, -aceae, etc.) suggest is relative inclusion relationships of groups of species in the local hierarchy. If we are talking about a monophyletic group, say Ericaceae, and Vaccinioideae are mentioned, then the latter must refer to a clade contained within the former, but neither is necessarily comparable with Polemoniaceae and Cobaeoideae, or any other family-subfamily combination. All are monophyletic groups (I hope - but see below), but that is all they have in common. Taxa at the same rank are equivalent only by designation and have nothing necessarily in common other than - perhaps - their monophyly. Rank as used here has no meaning other than signifying a monophyletic group that includes other monophyletic groups with appropriately subordinate rank terminations. Even sister taxa have only age and their immediate common ancestor and its ancestors necessarily in common. The non-equivalence of taxa at the same rank has long been clear (Darwin 1859; see also e.g. Seberg 1989; Stevens 1997; Bertrand et al. 2006; Giribet et al. 2015 and references), although they have sometimes been compared (Ricklefs & Renner 1994), if almost unconsciously, as in comments like "Orchidaceae are a very speciose family" - although I discuss here how even this apparanelt innocuous statement is in fact rather problematic.
A flagged hierarchy is useful in memory and communication (e.g. Stevens 2006a). It improves memorization by tapping in to the hierarchical structure of language (an extension of the noun-adjective structure of binomials); emphasis on genera, families and orders, as here, is simply a didactic device. Most genera, families, orders, etc., are monophyletic and for the most part include largish groups of species, otherwise they are ultimately just units that are useful in communication and that are used in general conversation by biologists and others world-wide.
However, Valentine and May (1996) described the Linnaean hierarchy as an example of an aggregative hierarchy with emergent properties at the different levels, while they dismiss phylogenies as being simply positional structures lacking emergent properties (for hierarchies, see also Eldredge 1985; Salthe 1985). Indeed, in the past rank terminations did denote absolute rank, the classification of Linnaeus (at least in theory) at the level of genus and species being such an example. Classifications where rank is absolute, taxa at the same rank being legitimately comparable entities and with particular (kinds of) properties, are class hierarchies in the strict sense (Stevens 2002, 2006a).
It is sometimes suggested that taxa at the same rank should have a similar level or number of morphological and/or molecular similarities or differences, or be based on similar characters, or be similar in these respects to taxa elsewhere in a larger clade under consideration (Heenan & Smissen 2013 for an example). This will lead to name instability if used in taxa where such criteria had not previously been used, especially if attempts are then made to extend such criteria beyond the local group of interest, and they may also imply that taxa at the same rank are equivalent and comparable in a biological context (for an example, see Fritsch et al. 2008). Of course, equivalencies in morphological and/or molecular differences are unlikely to be fixed (new apomorphies will continue to be found, or what were thought to be apomorphies turn out not to be) or easily quantifiable (especially with molecular data coming from different genomes). It has also been suggested that rank could reflect the age of the clade (e.g. Hennig 1966; Holt & Jønsson 2014; Eguchi & Tamura 2016). However, to say that dating clades is still a difficult enterprise is very much an understatement (see below), furthermore, if age were introduced as a ranking criterion, huge - if arguably only temporary - disruptions to the names that we use would result, certainly if this criterion were used across all of life. In this context, some recent proposals which invoke the use of age in classifications focus more on providing a standardized timeclip, i.e. set of letters referring to a particular geological period, that could be added to a conventional taxon name (Avise & Mitchell 2007; Avise & Liu 2011; see also Vences et al. 2013).
Indeed, taxa at the same rank are unfortunately still sometimes treated as if they were equivalent by those attempting to understand evolutionary or biogeographic problems (e.g. Ricotta et al. 2012), and some still worry about the properties of genera (e.g. Strand & Panova 2014); Hendricks et al. (2014) discussed the (mis)use of the generic rank by palaeontologists. Interestingly, in the middle of the last century, at least, taxonomists thought that the rank of genus was more "real" or "natural" than that of species (Anderson 1940), but a recent (rather small) survey suggests that opinions have flipped, with most thinking species are more "real" (Barraclough & Humphreys 2015), although discussions about "reality" and "nature"/"natural" are somewhat passé. However, when thinking about species and speciation, note that the general lineage species concept (de Queiroz 1998, 2007) and its descendants make a clear distinction between species concepts and the criteria that can be used to recognise species in connection with particular concepts. Indeed, coupling the uncertainty over what might be an individual (e.g. an apomictic dandelion; different parts of an ancient clone that now have no organic connection; the microbiome associated with a plant), and the availability of massive amounts of data (e.g. Novikova et al. 2016: Arabidopsis and relatives), means that deciding on the limits of species, let alone applying concepts such as monophyly to them, is becoming increasingly difficult (see also e.g. Freudenstein et al. 2016b; Zachos 2016; Barraclough 2019; Karbstein et al. 2024: the future? for general discussions about species). Maddison and Whitton (2023) present cogent arguments that there is no such thing as a species rank, and certainly one should not get too excited about estimates of species numbers mentioned here.
Understanding the distinction between grouping and ranking is important. We can both agree that there is a genus Acer, yet disagree as to whether it should be in Aceraceae or submerged in Sapindaceae. Although from one point of view such disagreements are utterly trivial, they still exercise the botanical community, perhaps because this is more a matter of personal preference than anything much to do with whatever science might be (e.g. c.f. Heenan & Smissen 2013 - splitting Nothofagus and Hill et al. 2015 - lumping it; Schuettpelz et al. 2018 - splitting ferns, and Christenhusz & Chase 2018 - lumping them; Ohashi et al. 2018 - splitting Desmodium).
Turning more specifically to phylogenetic classifications in general, and to the classification used here, Backlund and Bremer (1998) provide a very useful discussion on the principles of phylogenetic classification that is applicable at all levels apart from species. The zoologists Vences et al. (2013) list classificatory principles that are largely in agreement with those of Backlund and Bremer (1996) and those followed in this site, although Vences et al. are less concerned about having small genera that I am, perhaps partly because the groups on which they focus are small compared to those under discussion here. See also Stevens (1998), Albach et al. (2004), Entwisle and Weston (2005), Pfeil and Crisp (2005), Pauly et al. (2009), Bateman (2012 and references), The Legume Phylogeny Working Group (2013b), Y. Tang et al. (2015: pp. 22-24), Giribet et al. (2016), Kadereit (2017), etc., and many other papers are mentioned in the "Classification" section at the end of each family discussion.
The discussions in these and other papers can be applied to supraspecific classifications in general. Whereever possible taxa that are recognised formally should be monophyletic (but see below!). However, this does not indicate which particular clades we might wish to name as families, genera, etc., and so to talk about in general conversation: If a well-supported hypothesis of monophyly is a necessary prerequisite for a group to be named, it is not a sufficient prerequisite (but c.f. the PhyloCode - Cantino & de Queiroz 2006). Additional criteria should be invoked. Other things being equal, it is helpful if 1, taxa recognised formally are easily recognizable, 2, groups that are well-established in the literature are preserved, 3, the size of groups is taken into account, numerous small groups having little to recommend them since individually they summarise little information and tend to clog the memory, while groups that are too big may be amorphous, and 4, nomenclatural changes, e.g. if a particular clade is recognized as a genus, are minimized. These are the accessory criteria of Backlund and Bremer (1998) and which remain useful guidelines. The broader phylogenetic context should also be taken into account, that is, formal recognition of a taxon may necessitate the recognition of additional taxa at that rank if, for example, there are several pectinations immediately below the taxon of immediate interest since the former will also have to recognized at the same rank (e.g. Stevens 1998). Examples are the three or four small families that would be needed in Dipsacales if Valerianaceae and Dipsacaceae are maintained (see the broadly-drawn Caprifoliaceae) and the proliferation of genera caused by the piece-meal dismemberment of Spermacoce (Rubiaceae) or Desmodium (Fabaceae). It is essential to understand the overall phylogeny of a group before changing names in part of it (Bateman 2012; Guo et al. 2022), and this should be considered another accessory criterion; Lesica and Lavin (2023) discuss potential nomenclatural instability in the context of phylogenetic work.

Thinking of aspects of number and size of groups to be recognized, findings in ethnobiology and cognitive psychology can be used to suggest that a moderate number - probably fewer than 500 - of families is a reasonable goal at which to aim, and that groupings of taxa throughout any system should be rather small in size (e.g. Berlin 1992; Stevens 1994, 1997). A paper by Miller (1956), 'The Magical Number Seven, Plus or Minus Two: Some Limits on Our Capacity for Processing Information' (it had been cited 41,013 times as of 8.viii.2023) explains what is going on. Major systems such as those of Linnaeus and of Bentham and Hooker were constructed explicitly to ease the burden on the memory, the latter in particular ensuring that all groups in their classification were relatively small, often containing three to eight immediately subordinate taxa - but by no means all their groups were formally named (Stevens 1997, 2002; see also Scharf 2007; c.f. in part Vences et al. 2013). Along the same lines, Burtt (1977b) suggested that the number of names at any rank should be at most one third of those at the immediately lower rank - and monotypic taxa might not need formal names (Burtt 1977b) - and certainly to have monotypic subfamilies, tribes and subtribes for the one species is useless (Amborella-Amborellaceae and Humbertia-Convolvulaceae are just two of many places where one could do this). However, as phylogenies become more stable, it will be clear that "the shape of the tree" and the limitations of the Linnean hierarchy is resulting in some ugly classifications...
The accessory principles of Backlund and Bremer (1998) should be used in combination. Thus keeping the monogeneric Platanaceae separate from their sister taxon, Proteaceae, is justifiable: Both are much-used names that signal well supported, well defined and easily recognisable groups that have long been recognised as distinct, and both have several synapomorphies and do indeed look very unlike each other. Combining the two because Platanaceae are small (one genus, ca ten species) would yield a clade with few obvious apomorphies, not to mention the fact that Nelumbonaceae should by the same logic (it is also monogeneric, and with only two species) also be included in the expanded family... On the other hand, it is difficult to justify the continued recognition of Callitrichaceae or Hippuridaceae, monophyletic and distinctive although they may be. If they were recognised, several poorly characterised clades would also have to be carved out of Plantaginaceae in any classification that aimed to convey a comprehensive view of the world's flora (see also Pfeil & Crisp 2005; Orthia et al. 2005; Albach 2008, etc., for good practical discussions of such matters). But there are no absolute guidelines. Podostemaceae are sister to Hypericaceae, previously in Clusiaceae s.l.. The dismemberment of Clusiaceae s.l. allows the continued recognition of the very distinctive and long-established Podostemaceae, and Hypericaceae, Calophyllaceae and Clusiaceae s. str. are all characterizable and of moderate size; the recognition of Rafflesiaceae is a similar example. A somewhat similar situation: If Lemna and its relatives are a clade sister to most other Araceae, "should" they be recognised as a separate family? Gymnostachys, a phenetically fairly distinctive taxon, as well as the less phenetically distinct Orontioideae would also have to be recognised as a separate families, too, and Araceae in a somewhat more restricted sense would indeed be morphologically more coherent, although not greatly so, yet not so notably distinct... Unfortunately, it is in the very nature of such decisions that they seem ultimately somewhat arbitrary and unsatisfactory.
Takhtajan (1997) suggested that smaller families are more "natural" than larger families. This is incorrect. Although "having more features in common" is one common meaning of "more natural" (see below for more on "nature" and "natural"), monophyletic groups that include fewer taxa - Takhtajan's smaller families, for example - do not necessarily have more apomorphies than larger groups. A group of any size may have apomorphies, and any inverse correlation between size of group and apomorphy number is at best weak - think of all the apomorphies of angiosperms and monocots. However, unreversed apomorphies are more common in smaller, less inclusive clades. But by this kind of argument all families should be very small, since then their members will have a great deal in common, and so will be most "natural".
The slippery slope is obvious. As families (for example) are split, the relationships evident between the segregates and that were responsible for their being placed in a single family in the first place will seem to necessitate the recognition of a new order, etc., as is evident in Takhtajan's own work - general taxonomic inflation is the result (see also comparable suggestions in a cladistic context for Brassicales in particular - Ronse de Craene & Haston 2006). Such splitting is also questionable when teaching and learning families, since the student needs to understand the system as a whole. Nevertheless, for some genera removed from the families that until now have included them, the phenetic-classificatory-phylogenetic structure in their new home may mandate the recognition of small families. Note that Takhtajan's suggestion that narrowly defined families (better: taxa in general) are more useful for phylogenetic studies may be true, but this is a separate issue. Indeed, I have more than once regretted prematurely combining groups, whether species (in the context of monographic work) or families (in the course of preparing these notes).
To summarize: The emphasis here is on establishing a consensus classification, and on thinking of classifications as simply being means to an end, not an end in themselves. It is the phylogeny that is central, and the names in classifications are simply names attached to larger or smaller branches of the phylogenetic tree that facilitate our discussion about that phylogeny. A prerequisite for developing such a consensus classification is a stable phylogeny, and indeed, over the last 20-30 years family circumscriptions have been largely stable. However, relationships within orders like Malpighiales and Ericales remain unclear, and those between orders at the base of the rosids and asterids in particular are also uncertain, as recent work involving nuclear genomes is emphasizing. Indeed, the Plant and Fungal Tree of Life (PAFTOL) project seems to have got off the ground, and angiosperm phylogenies with much improved generic sampling and various kinds of analyses of nuclear data (W. J. Baker et al. 2021a, b) are increasingly frequent - the "Seed Plant Tree of Life with 10,709 angiosperm terminals (8336 genera - as of 10.x.2023) gives one view of the findings (see also Zuntini et al. 2024).
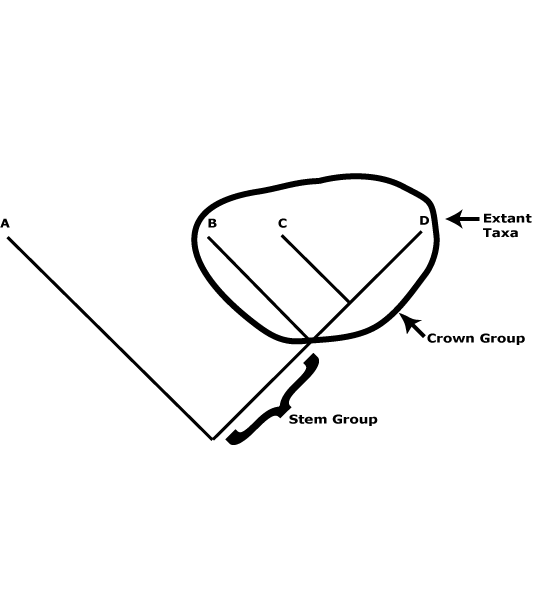
There is a useful distinction to be made between crown and stem groups. The former are the groups that include the extant members of a particular clade and their immediate common ancestor (see the figure below), and also any fossils that can be placed in the crown group. The groups characterized in this site are such groups. Thus Proteaceae are crown-group Proteaceae, apomorphies like the single carpel, four-merous perianth, etc., being found in their immediate common ancestor. Stem groups, on the other hand, include all the members of that clade below the crown group to immediately after its split from its sister group - and all branches (which have fossil representatives only, of course) of that part of the tree. Thus stem Proteaceae would include everything after the split from its sister group, Platanaceae, but they might well be unrecognizable as the Proteaceae of these pages. Obviously, most of the organisms in that part of the tree are unknown, only a few fossils being placed there, and it is also not known when/where particular apomorphies of crown group Proteaceae evolved along this branch. In the case of the stem group of angiosperms, not only is it largely unknown and probably well over 100 m.y.o., but almost certainly most of the organisms to be placed along it will be gymnospermous. By and large, we know very little of stem groups in general.
Consistent with such ideas, a fairly broad view of families and orders is taken here, as in the A.P.G. system, whenever the constraints of a rather relaxed view of monophyly and other criteria used when constructing classifications permit. Indeed, there is a principle in evolutionary classification that is quite useful here: The size of the gap between two groups tends to be inversely proportional to the sizes of the groups involved (Davis & Heywood 1963). In some situations a large group is formally divided even although the distinguishing characters of the two parts are weak, whereas a smaller group that is similarly divisable may be left intact.
There are ultimately no reasons other than convention or convenience why any group should not be segregated into several smaller groups, or merged to produce a larger unit; we can talk about one large thing, or about several smaller things. Here I follow van Steenis (1978), Philipson (1987b), and others who have questioned the utility of splitting a group when ideas of the relationships of its constituent members have not changed - that is, very good reasons have to be provided for splitting a family if the genera within it remain part of the same clade, rather than belonging to another clade. Thus A.P.G. (2003) broadened the circumscription of Malvaceae because of the para/polyphyly of some of the families that had historically been associated with it (Judd & Manchester 1997; Alverson et al. 1999; Bayer et al. 1999). These families, particularly Tiliaceae and Sterculiaceae, were not at all easy to distinguish, their close relationships had long been recognized (see e.g. Brown 1814), molecular studies were finding extensive paraphyly, and to some workers, at least, their combination was something of a relief. Although most of the larger clades within Malvaceae s.l. remain difficult to distinguish, even with flowers, Cheek (2007) opted for their wholesale and novel dismemberment into ten families; the "very good reasons" for doing this are wanting.
The same principles are of course applicable when it comes to dividing genera. Groups in which there is substantial disagreement over taxon circumscription despite basic agreement over classificatory philosophy include Orchidaceae, Asparagaceae-Scilloideae, Rubiaceae-Spermacoceae. However, little other than a headache is gained by splitting genera such as Ceratophyllum, Drosera and Gnetum as has been proposed (e.g. Doweld 2000). Indeed, even if there is well supported phylogenetic structure within a genus, this is certainly no signal that two or more genera have to be recognized, even if the new clades are consistent with morphology and had been recognized as genera some time in the past. Thus I think that the dismemberment of Pterostylis (Jones & Clements 2002b) was somewhat unfortunate (see Janes & Duretto 2010 and Schuitema & Adama 2011 for something close to a return to the status quo ante); that a previously unnamed clade is morphologically distinct and has synapomorphies does not necessitate its formal recognition (c.f. Clements et al. 2011). Along the same lines, if a newly-discovered taxon is sister to an existing named genus, this does not mean that a separate genus is needed for the newly described species (c.f. Davis 2002; Léveillé-Bourret et al. 2017). Humphreys and Linder (2009) provide a well-documented survey of generic concepts in plants which the reader should consult; they note that generic limits (broad versus narrow) have oscillated historically, and that currently larger genera tend to be recognised because studies tend to be on a broader scale than in the past - and for the record, broad limits are my preference, too.
Note that invoking "similarity" or "difference" - whether qualified ("considerable similarities", "substantial differences") or not - in a cladistic context as justification for combining or splitting taxa is not a particularly strong argument (see e.g. Cardiopteridaceae/Stemonuraceae - Kårehed 2002c). Similarity and difference cannot be defined precisely, since what may seem to be substantial similarities to me, may not to the next person, nor are they likely to be stable, our knowledge of morphology and what might be synapomorphies being in a state of flux.
I might have prefered to merge some families recognised here or split others, but by and large I do not think my own preferences matter very much - and I take the same position with regards to comparable preferences expressed by others. Indeed, the bottom line is that in flagged hierarchies of the kind used here, the limits of any monophyletic unit generally taught and discussed can be established only by convention and consensus (e.g. Stevens 2002, 2006a; Entwisle & Weston 2005; James & Duretto 2010). Given the increasing molecular and morphological support for the outlines of angiosperm phylogeny detailed in these pages, a stable classification based on this phylogeny seems attainable (e.g. Reveal 2011). Indeed, in addition to providing current ideas of relationships of seed plants in a synthesised form, this site is part of an attempt to build such a consensus over taxon circumscription (see A.P.G. 1999, 2003, 2009, 2016; Grass Phylogeny Working Group 2001, 2010; Mabberley 2017: also Hibbett et al. 2007 for a good example in fungi and Boyle et al. 2013 for a useful name resolution service).
Reaching a consensus is important, since what we know of angiosperm phylogeny allows a very large number of classifications to be based on it, and unfortunately, "nature" does not dictate what the classification should be, despite our almost universal protestations that our classifications are "natural". All classifications are constructed by humans to communicate particular aspects of groups, their characters, and relationships; they are means to an end, not an end in themselves - in this context, the papers by Bob O'Hara (e.g. 1988, 1992, 1993) are still very much to the point. Our goals as systematists are surely to produce robust hypotheses of relationships, to understand the evolution of morphology, and the like - having a tree allows one to think about evolution of evolutionary novelties or apomorphies, and how or even why homologies change over time, in a way that was previously very largely impossible (O'Hara 1997; Wagner 2015). However, to argue ad nauseam whether something "should" be a family or a subfamily, or whether a genus, known to be monophyletic, should be split, or a group of genera, all monophyletic, should be merged, is largely a waste of time. It is not simply that it is difficult to read literature in which different taxon limits for the same plants are being used, or different names used for the same clade, but the unintended consequences can be considerable. Thus the acceptance of generic name changes in some Orchidaceae-Epidendroideae in Pridgeon et al. (2005) meant over 10,000 changes to the names of orchid hybrids which had to be transferred to a hybrid genus other than the one in which they were originally registered (Royal Horticultural Society 2008) - note that here something did have to be done about generic limits. Changing names unnecessarily surely leads to madness, and worse: The discredit of our discipline.
There are similar issues whatever naming system is used. Thus in phylogenetic naming (Baum et al. 1998 for an example, but c. f. Baum et al. 2004; for the PhyloCode, see Cantino & De Queiroz 2006 and especially de Queiroz & Cantino 2020; for examples of a number of PhyloCode-type names, see de Queiroz et al. 2020) an unflagged hierarchy is used in which terminations of names used are uninformative about the relative position of taxa. Such unflagged hierarchies have serious deficiences as communication devices. They lack one aspect integral to classifications, whether of vehicles, of organisms by local people, or of organisms in the context of a phylogeny - they contain no intrinsic information about the relationships of the group in question to any others (e.g. Pfeil & Crisp 2005; Stevens 2006a). Recent suggestions for using prefixes like "Apo-" and "Pan-" to PhyloCode names might, however, allow limited information of this kind to be conveyed, but only as it pertains to individual branches, while other dodges that may help include the use of prefixes and suffixes like Holo-, -formae, -morpha, Eo-, Eu-, and Neo- (e.g. Penagos Zuluaga et al. 2021). In any event, such proposals simply prevent the tripling (or substantially more) of the number of quite different names that could be used to describe different aspects of the one phylogenetic tree. In general, where n is the number of extant species in a group, the number of clades in such a group = n-1. (Species will also need names, too; for their names, see e.g. Dayrat et al. 2008; current PhyloCode thought is to leave the naming of species alone for the time being. However, for practitioners of the phylocode the species epithet is the only part of the name that must be used.) Of course, lumping and splitting mercifully have no parallels in phylogenetic naming, the circumscription of clades associated with names remaining largely invariant, and old Xaceae s.l. and Xaceae s.s. names will be replaced by separate names; any well-supported clade can be named without affecting the name of more or less inclusive clades. Indeed, the hierarchy of the conventional codes of nomenclature simply cannot handle naming all the clades that one may wish to talk about when discussing phylogenies. How the PhyloCode will deal with reticulation is unclear.
Consensus over the clade names commonly learned by students, used in herbaria, and, importantly, used by the general biological community will still be needed, otherwise communication will be impeded. Alternative classifications are an impediment both in bioinformatics and in general communication (e.g. Vorontsova et al. 2015). PhyloCode names in themselves contain no guidelines as to which names should be chosen. But the situation is still more complicated. Terminations that convey ideas of rank in a phylogenetic classification can also be used as PhyloCode names - the use of such names is neither encouraged nor discouraged - however, there they will carry no implications of rank. How they will be used is another matter.
Of course, there are other ways of constructing classifications, and some still prefer evolutionary classifications (see e.g. Ann. Missouri Bot Gard. 100(1-2). 2014 - somewhat dated, c.f. Schmidt-Lebuhn 2010; Grímsson et al. 2017c and Bomfleur et al. 2017: fear of "ancestors"?). There classificatory principles differ substantially from those followed here, e.g. the recognition of paraphyletic taxa is allowed. Evolutionary classifications in general try and combine phylogeny and morphological gaps, although that is no easy thing to do - it is akin to combining chalk and cheese, the result being of little use either for blackboard work or for eating. For attempts to make this impossible task seem more objective, see Stuessy and König (2008) and in particular Willner et al. (2014), although from the comments made throughout this site it is clear that using such classifications as a source of apomorphies, as in Willner et al. (2011), is a burden they cannot bear. Some (e.g. Thorne 1976) suggested that the sizes of gaps between groups at the same rank should be similar, but any principle like this is inherently flawed (see also above). The basic problem with paraphyletic groups is that users do not generally expect a family to be nested inside another family, a genus within another genus, etc. This flouts the basic principles of classifications in general as evident in folk taxonomies and the general expectations of language. Thus when I was working on Ericaceae in the early 1970s it was thought that they were very uncommon in Australia, although it was noted that the related Epacridaceae were very common there, and this caused some bemusement; why were there so few Ericaceae in Australia? Given that Epacridaceae are now nested well within Ericaceae (see Epacridoideae), the largely language-driven biogeographic problem has vanished.
Systematists of all persuasions may invoke ideas of "nature" and of "natural groups" when delimiting groups (e.g. Stuessy & König 2008; Stuessy 2010). Unfortunately, the ultimate meaning of "natural" has long been "a group of the kind [usually unspecified] that I think should be recognised", so any classification competing with your preferred classification becomes ex ipso facto unnatural (e.g. Bather 1927). Its use is not helpful since one has first to find out what its user means; in general, the invocation of "nature" or "natural" in such contexts tends to preclude constructive discussion. I prefer not to use the word.
A number of new and largely independent classificatory systems have been coming out, including those by Takhtajan (1999, 2009), Doweld (2001), Wu et al. (2002), Goldberg (2003), Shipunov (2005 et seq.), Thorne (2007; very elaborate), Heywood et al. (2007), Reveal (2012: also elaborate, no justification), Willner et al. (2014), D. L. Fu (2019: different classificatory principles, well over 100 new family names, but all nomina nuda), and so on. However, even those who allow or promote the recognition of paraphyletic groups (e.g. Grant 2003; Thorne 2007; Heywood et al. 2007) may find a system recognizing only monophyletic groups of some interest - it can provide a rather different understanding of evolution.
The bottom line is that with a stable hypothesis of phylogeny, a classification based on that phylogeny can (must) be stable, too. We can then get on with our work, that is, testing the structure of phylogenies we have, elucidating phylogenies in areas where relationships remain unclear, studying the evolution of morphology, understanding eco-physiological changes, describing species, etc.. In this context, the spread of the Angiosperm Phylogeny Group system (see below) and its widespread use in technical literature, also floras (e.g. van der Meijden 2005), dictionaries (Mabberley 2017), textbooks (e.g. Judd et al. 2016), more general and popular literature (e.g. Souza & Lorenzi 2012 - the third edition; Spears 2006; Hilger et al. 2010), and in herbaria (Haston et al. 2007, 2009) is gratifying. Returning to Godfray and Knapp's (2004) users of classifications who want a stable, informative and accessible classification that enables easy identification - essentially, they want cake with everything - these pages attempt to satisfy as many of their needs as possible, but without compromising the primary, if now somewhat tempered, classificatory principle, that of the monophyly of the groups recognized.
On this classification in particular.
It would be impossible even to think about a higher-level classification such as this without the advances in our understanding of relationships made by the phylogenetic analyses of molecular data carried out over the last thirty years or so. These are coupled with morphological studies (but see below) that are evaluated in this new phylogenetic context. For the dramatic changes in this area, see, for instance, the pessimistic attitude about orders in Davis and Heywood (1963: 107-108); "The most unsatisfactory taxon in Angiosperm classification", they were "indefinable", their circumscription was not fixed, etc.. Families, they thought, were likely to be the largest "natural" unit recognizable in flowering plants. Art Cronquist's magnum opus, his An Integrated System of Classification of Flowering Plants, came out in 1981 just about the time that molecular studies were becoming common. Of the 63 families in his Liliopsida or monocots, there have since been additions to and/or deletions from 17% of them, while of his 19 orders, only one, Zingiberales, is recognized today with the same circumscription. In his Magnoliopsida or dicots there have been additions to or deletions from 43% of his 242 families, while none of his 33 orders has the same circumscription today. Families like Liliaceae, Smilacaceae and Grossulariaceae have suffered the most, genera that he included in these families now being found in 10, 7, and 7 other places respectively. Obviously, although he was interested in evolution, how he formed groups and recognised relationships (evolutionary trends were important to him) were quite different from how this is done today, and he was well aware that families like those mentioned ended up as the resting places for genera of uncertain affinities because they were already so heterogenous that adding other genera made little difference to the family description. It is interesting that his groupings have fared rather better in monocots than in "dicots".
Here I very largely follow the most recent version of the Angiosperm Phylogeny Group classification (A.P.G. IV 2016). Any differences are not to be interpreted as differences in principle, simply that new phylogenies continue to be published and that this site is designed to provide an overview of current ideas of higher-level relationships of all seed plants. The Angiosperm Phylogeny Group classification is based on relationships evident in the numerous molecular studies that began to appear in the late 1980s, much of it based on analysis of sequences of chloroplast markers (see A.P.G. 1999 for the principles underlying the classification), and the major outlines of the trees used by A.P.G. II (2003) or even A.P.G. I (A.P.G. 1999) have not had to be changed. We will see what happens when analyses of nuclear genomes become common - see immediately below.
There have been only minor changes in the composition of the orders, even if the odd genus or even family is turning out to be seriously misplaced - examples are Hydatellaceae (from monocots-Poales to Nymphaeales: Saarela et al. 2007), Guamatelaceae (from Rosaceae to Crossosomatales: Oh & Potter 2006), and Perrottetia and Bhesa (from Celastraceae to Huerteales and Malpighiales respectively: Zhang & Simmons 2006). The main changes have been clarification of the relationships of unplaced campanulid families (Winkworth et al. 2008) and of genera placed in the old Icacinaceae (see e.g. Stull et al. 2015 and euasterids). Petrosaviales is also uncertain (H.-T. Li et al. 2019: Gitzendanner et al. 2018a, b).
However, note that relationships between orders and within many of the larger orders are still unclear in part, and as nuclear genes and genomes become used more often, there may well be changes in the very scaffolding of the A.P.G. tree (for intimations of what is in store, see Sun et al. 2014; O.T.P.T.I. 2019; W. J. Baker et al. 2021a, b: theSeed Plant Tree of Life; H.-T. Li et al. 2023; Zuntini et al. 2024; see also Pentapetalae). Confusion has perhaps been introduced by the results of a survey - largely of taxonomists, it seems - that suggest there could be changes to family-level circumscriptions even in the absence of changes of phylogenetic relationships (Christenhusz et al. 2015; c.f. Cole 2015), which would seem to fly in the face of common sense, at least if stability of names is of any interest. If it is not, then surely the Phylocode may seem a better way to go.
Phylogenies are largely shown as tree-like representations, although as discussed below it is becoming increasingly evident that parts of the land plant tree are more or less reticulating. Most trees have been more or less ladderized, which means showing the smaller (in terms of numbers of terminals) sister taxon first at each node, and the sequence in the text follows that along the top of these trees. (Since phylogenetic trees are like mobiles, the only fixed points being the nodes, there are innumerable other ways to construct a sequence - e.g. Hawthorne & Hughes 2008.) The result is that the trees here tend to be pectinate. When reading a taxonomic treatment or following a herbarium sequence, pectinations, interpreted carefully, have their value (see Haston et al. 2007, 2009). As one reads the terminals of a pectinate tree left to right, adjacent terminals will tend to be separated by a minimum number of apomorphies. Nymphaeales and Austrobaileyales are here adjacent on the tree, but they could be separated by hundreds of families in another arrangement without offending any relationships; if adjacent in a book or herbarium, then it is relatively easy to relate their apomorphic characters, but if separated by hundreds of pages, or two floors in a large building, then it is less easy to get anything immediate from the sequence. (Since all orders on this site are preceded by the apomorphies of all the nodes immediately below them in the embryophyte phylogeny, and because of the linkages that have been built in, this particular problem does not arise; it is possible to read off all the apomormorphies within the embryophytes of all terminals.)
The best herbarium sequence based on a particular phylogeny would maximize the number of taxa that are successive branches of the tree that are also consecutive in the sequence. Since specimens are generally filed under families, the outline of a new family sequence for arranging herbaria can now be suggested (e.g. Haston et al. 2007, 2009: note that numbers of species in a terminal ultimately governs the sequence; Waern et al. 2013). This will further help teaching and learning about plants. There are other cognitive and perceptual issues to think about when drawing trees (e.g. Novick & Catley 2007; Novick et al. 2012); thus wind-blown trees with all the lines oblique, as in the trees here, may unfortunately be less immediately comprehensible than erect trees with all lines either vertical or horizontal.
In cases where the Angiosperm Phylogeny Group (2003) suggested alternatives as to the limits of families, e.g. Papaveraceae in the broad sense or Papaveraceae, Pteridophyllaceae and Fumariaceae in the strict sense, Proteaceae in the broad sense or Proteaceae and Platanaceae in the strict sense, the choices made in A.P.G. III (2009) largely follow common usage, as in textbooks like Judd et al. (20015), Murrell and Gillespie (2021) and Simpson (2019), and particularly in the successive editions of Mabberley's The Plant Book(see Mabberley 2017), Mabberley (2008) in particular being an attempt to reflect a consensus, the result of taking the opinions of botanists at several meetings. A.P.G. III (2009) thus dispensed with alternative classifications, and reasons are given for the choices that are made there. For many the existence of alternative classifications simply confused, so reaching an agreement over which names/groupings to use when alternatives are possible seemed desirable.
A.P.G. IV (2016) and this site continue to attempt to reflect consensus and stability, in particular, they are guided by the important principle, that if relationships do not change, then names should not. Major textbooks and other general treatments remain largely in line (e.g. Judd et al. 2015; Soltis et al. 2017; Keller 2023). Some names or authors of names may be incorrect according to the latest findings of bibliographic sleuths, but I wait until activities in this largely inconsequential area stop before making changes - indeed, these are sometimes negated by subsequent findings. For a very useful compilation of suprageneric names, see Reveal (2000 onwards), for a welcome proposal to stabilize familial and generic (and specific) names, see G. F. Smith et al. (2017), and for superordinal names, see Chase and Reveal (2009).
As mentioned above, higher-level relationships in general, and the composition and still more the relationships of orders in particular, have long been rather difficult for systematists (e.g. Davis & Heywood 1963), and the limits of orders have been far more labile than those of families. The composition of clades like Apiales, Crossosomatales and Pandanales is decidedly unexpected, however, these higher level clades are generally accepted even in works with different classificatory philosophies (a good example is Heywood et al. 2007). However, within clades like Malpighiales, Asparagales, etc., attempts to find distinctive characters have largely failed (but see Endress & Matthews 2006a). But as with families, groupings suggested by molecular studies may be supported by morphological and/or chemical characters. Thus Crossosomatales are characterized by the anatomy of their seed coats, while distinctive chemical similarities between Pittosporaceae and their apparently rather different sister group, the Apiaceae/Araliaceae area, have long been known (Hegnauer 1969b for references). As our knowledge of morphology and chemistry improves we can hope for gradual improvements in clade characterisations and the detection of apomorphies at all levels.
The discovery of the relationships of parasitic and aquatic groups in particular have presented a challenge to systematists. Morphologically, some of these plants are so highly modified that interpretation of the plant body in conventional terms is difficult or even impossible. Thus parasitic groups such as Rafflesiaceae are hard to place since both the vegetative body and the flowers are changed almost beyond recognition (see Naumann et al. 2013: comprehensive study of parasitic taxa and their evolution). The same is true, although to a lesser extent, in many plants pollinated by wind, etc.; a variety of characters may be affected, and wind-pollinated plants have independently acquired similar modifications. Morphological change is also extensive in holomycoheterotrophic/sapromycotrophic (Leake 1994) and carnivorous groups. Again, in the latter there are limited number of morphologies (e.g. pitcher leaves, sticky glandular hairs) involved in the carnivorous habit, and there may even be convergence at the molecular level, as in the convergent evolution of digestive enzymes (Fukushima et al. 2017), while holomycoheterotrophic monocots from three orders clustered together in an analysis by Lam et al. (2018) because of long branch attraction in the molecular data.
In molecular analyses of parasitic groups, plastid gene sequences may be difficult or impossible to obtain, the chloroplast DNA in particular experiencing extensive gene loss. Gene loss seems to have occured in parallel (independently in the different groups), yet a core of functional genes remains, probably because of functional constraints (Delannoy et al. 2011). In parasitic plants the rate of molecular change in general is often high (e.g. Duff & Nickrent 1997; Nickrent et al. 1998; Caddick et al. 2002a; G. Petersen et al. 2006b; Barkman et al. 2007; Bromham et al. 2103), and the inclusion of parasitic taxa in molecular analyses can cause conniptions (e.g. Nickrent et al. 2004; Davis et al. 2004; Chase et al. 2006; G. Petersen et al. 2006b), not least because there can be horizontal transmission of genes (e.g. Davis & Wurdack 2005: Vitaceae to Rafflesiaceae; Barkman et al. 2007: the mitochondrial atp1 gene commonly moves) including the the invasive cox1 intron (Barkman et al. 2007).
Progress is being made. Thus placements for families like Rafflesiaceae, Mitrastemonaceae, Cytinaceae and Cynomoriaceae, all parasites and with highly modified morphologies, have been suggested (Barkman et al. 2004; Davis & Wurdack 2004; Nickrent et al. 2004; Davis et al. 2007; Bellot et al. 2016), and although it seems likely that Burmanniaceae s.l. are polyphyletic, here the parts are all to be located in Dioscoreales (Merckx et al. 2006, 2009a, 2010a). Similarly, plants groqing in the aquatic habitat are much modified - neither vessels in particular nor much xylem in general is needed; leaves are highly modified; and water-mediated pollination, if adopted, is usually associated with major changes in floral morphology. But here, too, molecular studies suggest that aquatic groups with hitherto problematic relationships may find homes. Thus Podostemaceae are close to Calophyllaceae and Hypericaceae (Malpighiales: e.g. Kita & Kato 2001; Ruhfel et al. 2011, 2013), Hydatellaceae, which used to be in Poales, are part of Nymphaeales (Saarela et al. 2007), and Hydrostachyaceae have a similar relationship with Cornales (Xiang et al. 2002; Schäferhoff et al. 2010; Xiang et al. 2011), although they have been placed in Lamiales (Burleigh et al. 2009). In the first two cases in particular there are morphological and chemical features that support the move. Very different placements of Ceratophyllaceae have been suggested over the years, unfortunately, its relationships still remain uncertain (e.g. Doyle et al. 2015; Kvacek et al. 2016 and references). If Ceratophyllaceae are sister to eudicots (e.g. Moore et al. 2007), they are so derived that there is no morphological evidence of which I am aware to support this relationship. Needless to say, as relationships for such wind-pollinated, parasitic, etc., taxa, are clarified, it changes one's understanding of evolution, both of the groups themselves (e.g. Hardy & Cook 2012) and the clades in which they find new homes.
In many cases the "new" family limits of the Angiosperm Phylogeny Group (see A.P.G. 1999, 2003, 2009, 2016) are not really controversial, although changes from the limits commonly accepted only a decade ago are sometimes quite dramatic (e.g. Wagenitz 1997). Thus the split of the old Saxifragaceae s.l. is necessitated by its extreme polyphyly, as also with Icacinaceae and Cornaceae. Although such groups had long been considered unsatisfactory, there had been no compelling evidence allowing the user to prefer one circumscription over another; now there is. It is generally accepted that the limits of Lamiaceae and Verbenaceae have to be redrawn, and the content of the two has changed considerably, however, they are now easier to identify than before. Hence the decision to recognise the recircumscribed families is not difficult, and if they were not kept separate, much of Lamiales would collapse into a single family. The same is true for Salicaceae and Achariaceae (Malpighiales), two previously small families that have received the bulk of the old Flacourtiaceae. Clade and hence taxon limits do remain difficult in some Malpighiales, but even there clade support is strengthening (Xi et al. 2012b). Euphorbiaceae have presented problems - in the past, one could recognise Euphorbiaceae s.l. quite easily because of their explosively-dehiscent fruits with a distinctive persistent columella. Putranjivaceae, one quite unrelated segregate, are quite distinct, as is confirmed by chemical evidence (presence of glucosinolates/mustard oils), but there are four families that make up the bulk of the old Euphorbiaceae that remain in the same part of the Malpighiales tree (e.g. Wurdack et al. 2004; Davis et al. 2005). However, there is no molecular evidence warranting combining just these Euphorbiaceae segregates, and if they were to come together, the clades in this part of the tree suggest novel groupings not recognised before. Thus Rafflesiaceae appear to be embedded within the single-ovulate ex-Euphorbiaceae (Davis et al. 2007), and there is little enthusiasm for reducing the iconic Rafflesiaceae to synonymy. In the sister clade are biovulate ex-Euphorbiaceae - and the well-known Linaceae. Relationships in core Caryophyllales, especially around Phytolaccaceae (perhaps somewhat surprisingly) and Molluginaceae (less surprisingly) were incompletely understood and refashioning of taxon limits has continued as relationships become more apparent (see e.g. Nyffeler & Eggli 2010; Christin et al. 2011a; Brockington et al. 2011; Y. Yang et al. 2015; Christenhusz et al. 2014). The old Icacinaceae have been broken up (see in particular Stull et al. 2015), however, clade limits in the old Olacaceae, mostly hemiparasites, remain unclear (Nickrent et al. 2010 and references).
Many families in these pages are polythetic at the morphological level, that is, they lack unique features characterizing ("defining") all and only members of that family. They can be recognised phenetically only by the unique combinations of characters that they possess. This is the result of evolution; the synapomorphy characterizing a lineage may be lost or modified beyond easy recognition in some of its members, or the synapomorphy may appear to be identical to a feature that has evolved in parallel in quite unrelated plants. (Taxa in evolutionary classifications are also often polythetic.) Indeed, some families recognized here now include substantial variation as phenetically distinct derived groups have been placed in their proper phylogenetic position - examples are the erstwhile Empetraceae, a wind-pollinated group, now included in Ericaceae, and the various derived, small-flowered aquatic and wind- or water-pollinated groups that are included in the overwhelmingly large-flowered and animal-pollinated Plantaginaceae.
In the past, problematic taxa tended to be assigned to groups using a few or even only a single character weighted particularly strongly on a priori grounds. The result was the recognition of conglomerate taxa such as Amentiferae (wind-pollinated), Rafflesiales (parasites), Nepenthales (carnivores), etc. (Cronquist 1981), all of which have been found be highly polyphyletic, and/or the segregation of families like the wind-pollinated Plantaginaceae s. str. (now much expanded) and Leitneriaceae (now in Simaroubaceae). Given the ever-increasing amount and complexity of morphological knowledge and the difficulty of carrying out complex multivariate analyses in one's head, a solution in the days before computers was to emphasize just one or a few characters thought to be evolutionarily important and delimit groups accordingly. Thus the Parietales included a number of families all of which had parietal placentation - and these families are now widely dispersed on the tree.
Indeed, we used to assume that features like highly scalariform vessel perforation plates or complete absence of vessels, or a flower with an androecium that had many stamens, a superior ovary, or separate petals, were necessarily "primitive", and conversely vessels with simple perforation plates, an androecium with few stamens, inferior ovary, or petals that were connnate were almost necessarily "advanced" (but c.f. Stebbins 1951), or that variation in such characters was necessarily of evolutionary or taxonomic importance. Such assumptions are incorrect (e.g. Seberg 1989: chromosomes; Soltis et al. 2005b); for the evolution of vessels in asterids, highly parallel/with reversals, see Lens et al. (2016). Indeed, as one reads the literature, one can see how the interest in particular characters or kinds of characters waxes and wanes over time. Carpels may become secondarily free; carpels may fail to close, the seeds then developing outside the confines of the carpel, as in some Aspagaraceae-Nolinoideae, Violaceae, Berberidaceae, Malvaceae-Sterculioideae, etc.; in Peliosanthes teta, perhaps the only species in Peliosanthes (Asparagaceae-Nolinoideae) the ovary varies from superior to inferior (Jessop 1976: species limits here need investigation), as in Saxifragaceae, Apiales (esp. Pittosporaceae), Asterales (see Menyanthaceae), Poales, etc.; many-seeded carpels can evolve from few-seeded carpels (Razafimandimbison et al. 2008, but c.f. in part Beaulieu & Donoghue 2013); monoecy may be derived from dioecy (Schaefer & Renner 2010 and references) - and on it goes. Classic studies such as those by Babcock (e.g. 1947) on Crepis that assumed that evolution - in this case of the karyotype in particular - was unidirectional have needed comprehensive re-evaluation (Enke & Gemeinholzer 2008). Most if not all characters have reversed and/or evolved in parallel, even at the level of amino acid substitution, as in the independent acquisition of the phosphoenolpyruvate carboxylase (pepC) gene in C4 photosynthesis in grasses (Christin et al. 2007b; see also Bläsing et al. 2000; Anke et al. 2004; Reimann et al. 2004; Maia et al. 2012; Langel et al. 2010; Irmer et al. 2015; Fukushima et al. 2017; Later et al. 2018; etc.).
Given the suggested relationships of some parasitic and aquatic groups, it can be very difficult to understand how they have evolved from their more morphologically conventional relatives, however, it may well be that our preconceptions as to likely or possible evolutionary change are at fault. If Podostemaceae are sister to Hypericaceae, I look forward to seeing hypotheses to explain how the dramatic changes in the vegetative body that have made Podostemaceae so problematic for generations of systematists took place. That conventional wisdom has trouble in understanding or explaining how the morphologies of such groups can be related is largely a problem with conventional wisdom. However, it is interesting that neither Podostemaceae nor Hydrostachyaceae, another very highly modified aquatic, group with established orders in the recent Angiosperms353 study of nuclear genomes (see W. J. Baker et al. 2021a); it will be interesting to see what happens as sampling in that project improves,
Similar problems affect generic circumscriptions. Thus emphasising the importance of floral characters that reflect pollination syndromes when drawing generic boundaries has all too often led to taxa that are highly unsatisfactory phylogenetically (see e.g. Acanthaceae, Bignoniaceae, Campanulaceae, Ericaceae, Melastomataceae, Orchidaceae, etc.) while over-reliance on characters of fruit and seed (see particularly Brassicaceae and Apiaceae) has also led to questionable generic limits. Again, the more general problem has been caused by the use of one or a very few characters that have been weighted a priori to structure classifications (see also García et al. 2009). Working out limits for genera is rightly becoming a major preoccupation as some stability is now evident in higher level phylogenies and taxa. Here the discussions about changes to generic limits in the German flora - which of course have implications far beyond Germany - that have been assembled in Kadereit et al. (2016) make fascinating reading. Again, there are tensions between those who are happy with narrowly-drawn genera, other prefer their genera to be more broadly drawn; some of the changes have already been made, some are definitely needed, but remain to be carried out, and there are yet others where the basic phylogenetic evidence is still incomplete. In some cases piecemeal changes are being proposed based on a local perspective, while in others waiting for the bigger picture to materialize seems the sensible thing to do.
In conclusion. Some families as delimited here may not be easy to recognize, but remember that detecting relationships - use whatever characters you can, even if they are not obvious - and naming a plant - focus on obvious characters that may or may not reflect relationships - are quite different problems. Taxa, although "natural", may not be readily recognizable, indeed, it was in exactly this context that Lamarck described the basic principles of writing dichotomous keys in 1778. Of course, Lamarck's idea of nature was very different from ours - there was some kind of continuum of form between living organisms, with no major gaps anywhere - and this meant that his genera (for example) might well not be sharply distinct from each other. Perhaps the best way of identifying plants at the family level is by well-made multiple access keys, as in Watson and Dallwitz (1992a onwards: family limits there may differ substantially from those adopted here). Such keys free users from the constraints of dichotomous keys in which particular characters are needed at each step of the identification process before there can be further progress. Instead, using whatever characters are evident on a specimen in whatever sequence is enormously useful in identification; when linked to illustrations, glossaries, etc., the power of this approach is enormous (see Dallwitz et al. 2000 [2006] for the principles underlying the construction and use of such keys). Nevertheless, dichotomous keys such as those of Hutchinson (1973), Franz Thonner (Geesink et al. 1981) and Cullen (2006) have their uses, although even in the last-named, despite its date of publication, the family limits accepted do not reflect much in the way of recent findings. Furthermore, as Mary Barkworth (pers. comm.) pointed out, the structuring of information in keys may mean that their users can acquire general knowledge about plants - although of course taxa coming out adjacent in keys may well not be at all related, a fact that is all too frequently forgotten. Identification aids are proliferating (e.g. Utteridge & Bramley 2014), and Wilf et al. (2021, see also Wilf et al. 2016a) describe a database of thousands of correctly identified cleared and x-rayed leaves - i.e. features of the venation, etc., are clearly visible - that can be used in identification; this will be particularly valuable in the identification of fossil leaves, but such databases also become a more general aid for identification.
Molecular data can be used to "bar-code" plants, that is, identify them by simply by analysing the sequences of one or two genes, these genes being widely sequenced so very extensive comparisons could be made. There was much interest in this approach for a while (Kress et al. 2005b; Kress 2017), although this interest has rather languished. However, every so often plants are collected whose identity is completely uncertain. Swaenepoel et al. (2020) used molecular data to place the remarkable Tiganophyton in Brassicales as a new family while W. W. Thomas et al (2021) also used molecular data to place specimens that had languished in herbaria for forty or so years in a new genus, Aenigmanu, in the poorly-known Picramnniales-Picramniaceae, another placement that the morphology of the specimens would never have suggested. However, in such situations one should be sure that the leaf one removes from the packet actually belongs to the specimen (Muñoz-Rodríguez et al. 2022, but c.f. de Almeida et al. 2023)...
(When identifying large numbers of plants, even more efficient than either style of identification, and certainly lots more fun, is sight identification. However, unless one has a photographic memory, one has to build up a good general knowledge of comparative plant morphology, and for most of us it is on this the ability to make accurate identifications depends. When faced with an unknown plant, I always look for leaf teeth and stipules and dots of various kinds in the lamina and check leaf insertion; smelling crushed leaves can also be helpful. The short paragraphs added after most families may help in confirming familial identifications. In this context, nodal anatomy can usually be checked using a razor and a hand lens.)
Numbers of taxa? For the little that it is worth, there are 4 orders and 13 families of gymnosperms characterised on these pages, and together they include some 82 genera and 947 species. For angiosperms, comparable figures are 57 orders, 446 families, 13,208 genera, and 261,750 species (of which monocots include 11 orders, 89 families, 2,759 genera and 52,760 species). Note, however, that higher mathematics was never my strong point; numbers, especially of genera and species, change daily so any figures mentioned are more or less seriously out of date - and anyway they are pretty meaningless. Even for species, which some might think smacked slightly more of reality than other taxa, estimates range as high as 422,000 (Govaerts 2001), with perhaps 10-20% more currently undescribed (Joppa et al. 2010), and new estimates keep on appearing (e.g. Freiberg et al. 2020). Indeed, estimates of numbers of species in groups that one would have thought were well known vary widely, two examples being the daffodil genus, Narcissus (estimates range from 15-81 species: Marques et al. 2017), and the bee orchid genus, Ophrys, where 10-354 species are estimates made within the last fiften years or so (e.g. Bateman et al. 2011a; Vereecken et al. 2011; Sedeek et al. 2014; Alibertis 2015: photographs; Cuypers et al. 2022, see also below). Recent general estimates also highlight the issue: Christenhusz and Byng (2016) estimate that there are 295,383 species of flowering plants, while 369,434 species is the number in Nic Lughadha et al. (2016), and see also Freiberg et al. (2020). Furthermore, Mallet (2013: p. 690) noted that "species counts over large areas of space and time [e.g., those in these pages] represent only a sketchy measure of biodiversity, a measure that owes more to taxonomic and metaphysical fashion than to science". Nevertheless, as emphasized here, families are useful in teaching, we as a community can ensure that their limits remain largely stable, and by concentrating on relatively few of the larger families one can gain some familiarity with much of the world's flora. Generic names are even more used, and again, a goal here should be to ensure stability in generic circumscriptions.
In this context, Moonlight et al. (2024) give estimates of the numbers of species (from the World Flora Online - Borsch et al. 2020) in the 86 large (more than 500 species) and 28 megadiverse (more than 1,000 species) genera of angiosperms, and their estimates are largely followed below. This work builds on that of Frodin (2004), and it is interesting that over the years there has been relatively little change in the genera involved, although some have been broken up because they are not monophyletic and so have left the list, while others have been added, including at least three that owe their presence there to the large number of apomictic microspecies that they contain. These large genera include around about 25% of all species of flowering plants, and the megadiverse genera alone some 13.6% (Moonlight et al. 2024). Infrageneric classifications can be tricky here, and Muñoz-Rodríguez et al. (2023) discuss large genera with a focus on their infrageneric classification in particular – they had been working on Ipomoea. They suggest that infrageneric groupings should be monophyletic, their phylogeny should be resolved, they should be diagnosable, all species in the genus should be placed somewhere or other, and nomenclatural stability should be maximized. (These criteria are applicable to phylogeny-based classifications in general.) They note that many of the monophyletic groups in Ipomoea cannot be diagnosed, so this would result in undiagnosable genera if Ipomoea were to be divided, and they suggest that an informal classification is currently best for a broadly circumscribed Ipomoea, which itself can at least be diagnosed.
ON ANCIENT HYBRIDS, INTROGRESSION, ETC. — AND PHYLOGENETIC TREES WITH GRAFTED BRANCHES
[Back to Top] (Note: This section is under (re)construction.)
Problems with an emphasis on strict monophyly - a clade is monophyletic if it includes all and only the descendents of a common ancestor - can perhaps be caused by reticulation events such as endosymbiosis (Gray & Archibald 2012 for literature), lateral/horizontal gene transfer (see e.g. Dunning Hotopp et al. 2007; Boto 2010; F.-W. Li et al. 2014) and hybridization, however, the first two are unlikely to be major confusing factors in embryophytes. The major endosymbiotic events that characterize the clade of which flowering plants are a part (and gave rise to chloroplasts and mitochondria - for the surprisingly complex evolution of the former, see Sibbald & Archibald 2020) are very ancient and of themselves cause few problems for the student of multicellular organisms, indeed, patterns of subsequent movement of genes from the mitochondrion and chloroplast to the nucleus are phylogenetically interesting. Thus there is substantial lateral movement of mitochondrial genes in particular in some host-parasite relationships such as Tetrastigma-Rafflesia (Xi et al. 2012a; Yoshida et al. 2010), via chloroplasts in grafts (Stegemann et al. 2012: exchange between Nicotiana), and in Poaceae just about any pollen grain can at least germinate on any grass stigmas (well, a bit of hyperbole), and zygotes can be formed in crosses between e.g. Poöideae and Panicoideae (Lausser et al. 2010; Kellogg 2015 and references). Here, too, there are no major problems, providing one is careful, since the genome in such cases seems to be overwhelmingly from one source; such transfers do, however, raise all sorts of interesting biological questions (see Richardson & Palmer 2007 for a summary). There are other wide but currently inexplicable transfers of mitochochondrial genes to plants like Amborella from its epiphytic associates (Rice et al. 2013 and references), and even transfers of nuclear genes (Vallenback et al. 2008). J. Ma et al. (2022) and others discuss horizontal gene transfer (= HGT), e.g. from fungi and viruses to streptophytes, a process that has been going on for a very long time and that can have important evolutionary consequences.
More conventional hybridization and introgression are, however, another issue. In pre-2023 versions of this site I wrote, "Genome duplications - hybridization and associated allopolyploidy is a common cause of them - are common, and are turning out to be a major driver of seed plant evolution. They have occurred at various times during the evolution of seed plants (e.g. Soltis et al. 2009; Litt 2013: summary; Y. Yang et al. 2015; Landis et al. 2018; see also below). Many of these duplications do not currently pose major problems for either phylogeny reconstruction or the adoption of monophyly as the major criterion for groups to be recognised formally in phylogenetic classifications, certainly for genera and deeper clades." This paragraph now largely misses the point when thinking about monophyly, hybridization and higher taxa.
Polyploidy/hybridization (note, although often connected, the two are not the same) have been much discussed for the last century or more, and for recent papers that also cover the older literature, see P. S. Soltis (2005), T. E. Wood et al. (2009), Whitney et al. (2010), P. S. Soltis and Soltis (2012), M. S. Barker et al. (2012, 2016b), D. E. Soltis et al. (2016), Doyle and Coate (2018), Rice et al. (2019), Moran et al. (2021: more zoological), Stull et al. (2023), papers in American J. Bot. 103(7). 2016, Ann. Bot. 120(2). 2017, and so on. It is clear that the effects of hybridization are pervasive. Of course, it has long been accepted that more or less extensive reticulation, hybridization and/or introgression is common between species. For instance, Blanc and Wolfe (2004a) found that in 9/14 model species they examined (largely herbs) there was evidence of palaeopolyploidy. In a number of genera - Medicago is just one example of many - relationships at the species level are turning out to be highly reticulating (Maureira-Butler et al. 2008). Plants that belong to the one species of Senecio and Elymus may have two or more independent hybrid origins from other species (Yan & Sun 2012), introgression is widespread, and so on. The result is that there are numerous problems if one tries to apply a concept of strict monophyly to species, and many - but not all - biologists suggest that this concept is indeed inappropriate here (e.g. de Queiroz 1998; Funk & Omland 2003; much of the discussion in Hörandl 2006; Hörandl & Stuessy 2010; Lavin & Pennington 2022). Hybridization was thought to be uncommon in the tropics, except perhaps between herbaceous taxa (e.g. Gentry 1982), but there, too, recent work suggests that it it is quite common (Schley et al. 2022: focus on the Neotropics). M. S. Barker (2015) suggested that some 24% of angiosperms were hybrids, split more or less evenly between auto- and allopolyploids.
It is ancient hybridizations (see Stull et al. 2023 for a definition - genetic, e.g. the hybrid lineage has subsequently diversified or undergone anagenesis, not temporal), events that took place 2-100 Ma or more, that are a serious problem for a site like this. Here I mention just a few examples: Marcussen et al. (2022) suggest that most of the sections in Viola subgenus Viola are the result of hybridization. Similarly, many genera in Poaceae are allopolyploids, and the genera are ultimately based on different genome combinations (Dewey 1984; Löve 1984; Brassac et al. 2012: Barkworth 2000 for a history of Triticeae classification; G. Petersen et al. 2006a). Extensive reticulation is reported within Danthonioideae (Pirie et al. 2009) and perhaps particularly in Bambusoideae (Triplett et al. 2011, 2014; Y.-Q. Zhang et al. 2019; Tong et al. 2020 and many other papers - see Poaceae); recent work on the latter subfamily suggests extensive and deep hybridization (). There are recent and historical hybrids in Brownea (Fabaceae: Schley et al. 2020), intergeneric hybrids in Annonaceae (X. Guo et al. 2018) and in Betulaceae (Z. Wang et al. 2022: Carpinus × Ostrya), and in Brassicaceae, the recently-described Microlepidieae (Joly et al. 2009; Mandáková et al. 2010, esp. 2017) and Shehbazieae (German & Friesen 2014) are the result of hybridizations between members of different tribes. Indeed, for some years now there has been evidence of ancient hybridization events in other groups that at the very least cause discordance between relationships suggested by different genomic compartments, as in Smedmark and Anderberg (2007: Sapotaceae), Fehrer et al. (2007: Asteraceae-Lactucoideae), Y. Liu et al. (2013: perhaps Crepidinae × Lactucinae), Pelser et al. (2008, 2010: Asteraceae-Asteroideae), L. E. Watson et al. 2020: intertribal hybridization in Asteroideae), Tripp et al. (2013b: Acanthoideae: Acantheae × Justicieae), Folk et al. (2018b: Heuchera area, Saxifragaceae), García et al. (2017: Amaryllidaceae-Hippeastreae) and Dong et al. (2022: Oleaceae). Dated at (269-)248(-215) Ma the allopolyploidy hybridization event that resulted in the Boreoselaginella clade in Selaginellaceae seems to be the earliest known in vascular plants, the exact nature of the earlier genome duplications like the ε and ζ events being unknown (Kang et al. 2024).
Looking at this issue from a different point of view, the length of time that clades containing hybridizing taxa have been separated, we find that some current hybridization in Viola is between members of clades that diverged ca 19 Ma (Marcussen et al. 2022). Hybridization in Ficus is turning out to be remarkably common, even occuring between members of clades that have been separated for perhaps 70 Ma (G. Wang et al. 2020). In Osmundaceae extant members of clades that diverged before the Jurassic can hybridize (Bomfleur et al. 2014b/2015), and other deep hybridization is known elsewhere in ferns (H.-M. Liu et al. 2020; Stull et al. 2023).
Ancient hybridization, then, is a general problem, and the reason that most of the examples just mentioned are at the tribal "level" or below is because it is easier to detect hybridization events that are more recent. But as Clarkson et al. (2017: p. 1001) noted, "Numerous whole-genome duplication (WGD) or polyploidisation events have been identified in the evolutionary history of flowering plants, leading to the now well established concept that all angiosperms are paleopolyploids". Indeed, Wendel (2015) suggested that the genome of Brassica had been multiplied some 288 times, O.T.P.T.I. (2019) suggested that there had been some 180 WGDs in basal angiosperms, and so on. WGDs are often hybridization events (see also Heslop-Harrison et al. 2022 for genome duplications and polyploidy). Hybridization is also common in groups other than flowering plants, in ferns, for example, but less so in most gymnosperms. Now that nuclear genomes are commonly being analyzed in various ways, the extent of such problems in land plants - very considerable - is becoming clearer (e.g. Folk et al. 2018a; Rose et al. 2021). However, how to recognize and then to interpret patterns in molecular variation between clades that separated up to two hundred million or more years ago, with the putative hybridization events not much younger, is a non-trivial issue (see Stull et al. 2023 for discussion) - differences between gene trees and species trees are just a part of the problem, and there is no infallible way of recognising an ancient hybridization event. As Ringelberg et al. (2022a: p. 33) nicely put it, “[W]hat fraction of genes supporting a clade should be used as a cut-off for delimiting taxa? To what extent does it matter if there are alternative topologies that are supported by a substantial fraction of genes, even if that number is lower than the number of genes that supports the 'main' topology and what are the classificatory implications when only a small fraction of genes is informative for certain relationships." (see also Stull et al. 2023)? It was thought that there had been hybridization between ancestors of the fabid/N-fixing clade and the malvid clade the descendents of which are now represented by the COM clade (Sun et al. 2014; see also the rosids), but this now seems not to be the case. However, WGDs - basically, polyploids of one sort or another - with their attendant complexities such as extensive genomic rearrangements, duplicate gene retention (hence the problem of paralogs), biased fractionations, housekeeping genes in particular tending to become single copy, etc., are turning out to be very widespread in vascular plants (e.g. Soltis & Soltis 2012; Guo et al. 2022; for WGDs, see also elsewhere. Simple hybridization/allopolyploidy is just the tip of the iceberg: it is just one extreme of introgression, a very common phenomenon in plants, where genes from one or more clades end up in another clade by repeated back-crossing and there more or less replacing the genes of this other clade (see Hibbins & Hahn 2022 for a review). It can be difficult to recognize introgression and hybridization, especially when they happened long ago, and incomplete lineage sorting, which one may think of as being a species-level problem, can further confuse the issue, posing problems at surprisingly deep levels (see also Guo et al. 2022; Stull et al. 2023).
Indeed, the chloroplast, nuclear and mitochondrial genomes (c.f. gene trees and species trees; cyto-nuclear discordance), and morphology may all suggest different relationships, and hybridization may be involved. Thus there are cases of hybridization in which organellar genomes become introgressed yet the nuclear genome is little or not at all affected (Folk et al. 2016), and in syngameons like those in Quercus there may be a more or less continuous level of gene flow between entities that otherwise would seem to be "good" species (e.g. Cannon & Petit 2019). In Rubiaceae phylogenies based on mitochondrial genome variation question the hitherto accepted three-subfamily classification of the family (Rydin et al. 2017). More generally, it should be remembered that classificatory changes in land plants up only a couple of years ago (as of 2020) were often dependent on phylogenies based on variation in a few chloroplast genes, perhaps with the odd nuclear or mitochondrial gene thrown in, which has turned out to be somewhat problematic (e.g. see Gonçalves et al. 2020b, 2021). The whole chloroplast genome can be thought of as an uniparental marker that is effectively a single, if very complex, gene, and the mitochondrial is also equivalent to a single gene. Warnings against reliance on uniparental markers are very much to the point (Marcussen et al. 2014; see also below).
In the early days of cladistics the distinction between monophyletic (a group made up of all and only the descendants of a common ancestor and with a synapomorphy/synapomorphies), paraphyletic (only, but not all descendants of a common ancestor and often characterized by a plesiomorphy/plesiomorphies), and polyphyletic, based on convergent characters (Hennig 1950, 1966; see also Farris 1974) seemed clear, and taxonomists since then have attempted to base their classifications on such monophyletic groups. However, as discussed above, reticulations are clearly widespread, and so the idea of using strict monophyly as the basis of classification becomes decidedly problematic. However, Hennig (e.g. 1966: p. 208), had realized that species of plants might hybridize, although he was inclined to question whether the hybridizing entities were true species... Quite recently, Wheeler (2014) noted that mono-/para-/polyphyly did not deal adequately with a universe in which there was reticulation, and he proposed the terms periphyletic/periphyly, epiphyletic/epiphyly (c.f. epiphylly!), the two pairs of terms dealing with the consequences of a single reticulating event, network loss and gain respectively) and anaphyletic/anaphyly, a situation in which there is both network gain and loss. If the clade that resulted from the hybridization event were to be recognised as a separate taxon at the same level as the remains of the two parental clades, these latter would be periphyletic and the hybrid clade would be polyphyletic. Of course, hybridization scenarios can be very complex and the examples given by Wheeler suggest recent events, not the ancient hybridizations that are the focus here. Thus in Bambusoideae hybridizations amy involve members of fairly old clades that have no extant diploid members (e.g. Triplett et al. 2014 and references), and elsewhere in the phylogeny one is dealing with with putative hybridization events that happened 100 Ma or more.
How is one to deal with well-supported ancient hybridization events here? - the discussion in Stull et al. (2023) is useful. The name of clades that are of hybrid origin like Ericales and Oleeae are prefixed with "×", and followed by the names of the clades from which the putative parents came - these latter names will be directly linked to where they occur in the tree if they are not close (as in ×Ericales, but not ×Oleeae), as will those of the parents to their hybrid offspring. The use of ×Ericales Dumortier, ×Oleeae Dumortier, etc., for the names of hybrid taxa is similar to what goes on at the generic level, e.g. ×Crataemespilus E. G. Camus (hybrid of Crataegus and Mespilus - note that +Crataegomespilus is a graft hybrid) - in all cases, the authors of the names are the original authors, whatever they thought about the nature of the taxon they described; this all follows the International Code of Botanical Nomenclature (I am gteful to Kanchi Gandhi for discussion here). But do not expect all the changes needed, especially in groups like Poaceae, to appear overnight.
Of course, the clade that results from a hybridization event may itself be strictly monophyletic; it may include all and only the descendants of a common ancestor that in this case happens to be a hybrid. Extant members of the crown groups of the clades that include the parents of the hybrid clade may themselves also be strictly monophyletic, however, it is in the stem groups of these parental clades that problems arise. As an example, Dong et al. (2022: Figs 8, esp. 9) describe a rather complex history of hybridization and introgression, not to mention incomplete lineage sorting as well, in Oleaceae, q.v.. Here I mention only that Dong et al. suggest that Oleeae are the result of an hybridization event between stem Forsythieae (the staminate parent) and a ghost lineage of stem Jasmineae (the carpellate parent), and this resulted in the tetraploid ×Oleeae. Within ×Oleeae there seems to have been hybridization/reciprocal introgression between stem Ligustrinae and stem [Fraxininae + Oleinae], as well as hybridization/introgression from stem Fraxininae to stem Oleinae (Dong et al. 2022), but there the clades involved have remained diploid. (Of course, in nearly all such cases ghost lineages will have been involved in the hybridizations, but sometimes they will seem to be more ghostly than others.) Interestingly, members of both Jasmineae and ×Oleeae, alone in Oleaceae, produce oleoside (a route 1b/1c iridoid), and a scenario here might be that the ability to synthesize this arose early in stem-group Jasmineae and thence moved to the ancestor of Oleeae. Of course, the stems immediately subtending crown-group nodes in the seed plant tree can be very long indeed, over 100 My, and in some cases exactly when along the stem a change occurs will be of interest, but this problem is hardly unique to ancient hybridization events. However, Fig. 1 in Stull et al. (2023) suggests some quite different if plausible scenarios. In the predominantly blue clade at the bottom any apomorphies of crown group members may have appeared over a protracted period, while in the predominantly yellow clade one can imagine both the subclades as having their own apomorphies, even although they are not much younger than the hybridization event.
If there is a substantial preponderance of nuclear genes from one of the parents of a hybridisation/introgression event in the descendants, it would seem reasonable to place these descendants in the same clade that includes that parent. Thus L. E. Watson et al. (2020) were looking at five tribes of Asteraceae-Asteroideae and found that gene movement from one tribe to another was less than 20%; there might be no need to indicate reticulation events in such cases - see also the quotation from Ringelberg et al. (2022a) above, examples in Stull et al. (2023), and so on. However, in all cases, clearly describing and justifying the reasons for naming decisions is essential, and in some cases of extensive hybridization there will be a need for concensus, as perhaps in Triticeae, where the different classifications make things very difficult for the poor user who has to disentangle an unnecessary classificatory mess (e.g. Card et al. 2014: p. 102).
But all this still really avoids the issue. That is, with hybridization one cannot depict relationships as a simple tree, rather, the best way to represent hybridizations graphically is as reticula, the "phylogenetic trees with grafted branches" of the title of this section (e.g. Poczai 2013; Lutteropp et al. 2021; Guo et al. 2022; Schley et al. 2022; Stull et al. 2023), and adequately conveying such situations to the reader is a goal of a site like this.
ON FORMING CLADE CHARACTERIZATIONS AND THINKING ABOUT APOMORPHIES
[Back to Top]
The organization of the information throughout is largely hierarchical, that is, character states constant or almost constant in higher groupings are not mentioned at lower levels (see also Stevens et al. 2004b; Kellogg 2015). This is in line with a phylogeny- or tree-based system, yet it has, perhaps ironically, long been seen as being an advantage of many so-called natural systems, even those that owe nothing to evolutionary ideas (e.g. Cesalpino 1583; Jussieu 1789). However, there is much to do to make this style of presentation fully effective, and given the prevalence of reticulations in vascular plant evolution, one's thoughts about character hierarchies have to be tempered somewhat. In particular, whether character states more or less constant in a group are synapomorphies for it often waits for further clarification of relationships both within the group and between that group and its immediate relatives. For example, most Annonaceae have stamens with distinctive prolongations of the connective, but Anaxagorea is sister to the rest of the family and such connectives may not be a synapomorphy of Annonaceae, nor may indehiscent fruits and the absence of staminodes (e.g. Scharaschkin & Doyle 2005, 2006). As more is found out about phylogenies, such examples multiply, so the dismemberment of the Icacinaceae s.l. (Stull et al. 2015) has meant that the characterisation of the large euasterid clade is being clarified, which has important implications when we think about euasterid diversification.
The character hierarchy was built up by first drawing up lengthy descriptions of families and then fitting the characters in the descriptions to molecular-based trees with rather conservative topologies. Many of the states of some characters at the base of the angiosperm tree are fairly obvious, hence the fairly lengthy characterisation (apomorphies and plesiomorphies definitely mixed!) for the angiosperms as a whole (but c.f. Sauquet et al. 2017). For some characters, I have worked up the tree, placing them as high as the evidence suggested. Otherwise, features in common to each clade, whether order, families within an order, or groups of orders, are those that are as far as is known common to all the family characterizations in that clade; they may also be synapomorphies (but see below), and are placed at the lowest level in the tree for which there is information available. For some features I have used both approaches, but confusion should be minimal. As relationships and our knowledge of the variation within characters improve, the top-down and bottom-up approaches will merge.
Trees based on morphology alone often have little support, and they may silently lead the user astray (e.g. Zhang et al. 1992). Nevertheless, the validity of an approach that fits morphological variation to a molecular tree may be questioned (but see e.g. Doyle & Endress 2011 and earlier papers). However, I think it rather unlikely that branches that are well supported in molecular analyses will be overturned by morphological data, and if they were, then one would reconsider exactly what the different data sets were suggesting. Analysis of morphological data alone provides support for many of the clades evident in molecule-only analyses, and in conjunction with molecular data may lead to increased support for clades (e.g. Hufford 1992; Doyle & Endress 2000), although in Nandi et al. (1998) adding morphological data reduces support for a number of critical clades, too, a not-uncommon phenomenon. Indeed, in none of these papers is the use of morphology without ambiguity. Thus I have been wary of putting much weight on clades that have only morphological support, but molecule-based clades with low bootstrap or jacknife values (esp. below 70%) or posterior probabilies (below 0.95) are also treated sceptically - indeed, that we still rely so much on variation in the chloroplast genome alone to construct phylogenies is worrisome, although things are changing. Other reasons for prefering to fit morphological variation to a tree are mentioned below.
Morphological and molecular data are very rarely in irreconcilably strong conflict. However, examples that tend in that direction are the relative positions of the Monimiaceae and Hernandiaceae (Laurales), and there have also been arguments about the position of Hanguanaceae (Commelinales [see below; now very likely] or Zingiberales?), and of Triplostegia (is it in Caprifoliaceae-Dipsacacoideae or -Valerianoideae? - see Dipsacoideae). Although trees based on analyses of morphological data alone rarely command much support, Weins et al. (2010 and references) is an example of the integration of morphological and molecular data, and of fossils and extant organisms. Indeed, the general congruence between morphological and molecular data is impressive and heartening, and many clades can be characterised morphologically. It seemed in 1998 that there were no unambiguous morphological synapomorphies for angiosperm orders (K. Bremer 2000), and this is still true if by "unambiguous" is meant "non-homoplasious". However, many orders can be characterised and have synapomorphies (see below), even if these are homoplasious and morphologically indistinguishable (at least at the current level of morphological and developmental observation) from synapomorphies elsewhere on the tree.
Identifying apomorphies is important because understanding the evolution of characters - and the organisms that bear them - is one of our major goals. For this, several preconditions must be met. One needs to have an accurate, robust phylogeny, one has to have examined the right taxa both from the point of view of morphology and molecules, one has to have coded the characters correctly (i.e., delimited states appropriately), one has to have used the right model of evolution when fitting the variation to the tree, and finally, and little discussed here (but see e.g. the notes on the diversification and evolution of angiosperms), one has to establish the right temporal context and to factor in other relevant aspects of the environment (see e.g. Omland 1999; Stevens 2006b).
On each order page it is easy to access the features associated with each node leading to the order in question, and putative synapomorphies for the order, families, etc., will be found in the page itself (I began to indicate synapomorphies only in xii.2011), although it is clear that one has to be very cautious when talking about character evolution. If the synapomorphies here are compared with those in particular studies cited (e.g. Turgeon et al. 2001; Bremer et al. 2001; Endress 2001; Albach et al. 2001a; Judd et al. 2003; Judd & Olmstead 2004; D. Soltis et al. 2005b; Zhang et al. 2006), differences may be found. Although such studies have been integrated into the characterisations as far as possible, there are several reasons why there may be these differences.
- Firstly, I may not have found all the information about a particular character, there may be disagreement over its interpretation (e.g. Austrobaileyales), or I have added unpublished information (e.g. Primulaceae and their relatives, nodal anatomy, etc.; Diapensiaceae, leaf ptyxis; and Peridiscaceae, Centroplacaceae, etc.).
- Secondly, the sampling of nearly all molecular studies is incomplete, although it is fast improving. Unfortunately, the sampling of the morphological and chemical characters the evolution of which we are interested in understanding is also often very poor, yet it is not improving fast. For many anatomical, chemical and embryological characters that are confidently said to characterise families and other groups, we all too often have no idea if those characters are applicable to the whole clade, or just to a subgroup within that clade. Thus Albach et al. (2001a, see also D. Soltis et al. 2005b) assign possession of iridoids to the base of the asterid I + II clades. However, this feature is placed higher up the tree here, partly because of topological uncertainties, but partly because in Lamiales (for example), the first four clades that are successively sister to the remaining Lamiales either lack iridoids or (most Oleaceae) have iridoids different from those found in the other members of the clade. Similar problems arise when thinking of the evolution of ellagic acid in Ericales (Stevens 2006b). Indeed, Much older literature, especially that from ca 1870-1920, although there are earlier classics like Hofmeister (1849), for example, remains invaluable, since it includes surveys, often never improved on since, of various aspects of plant anatomy and morphology for particular groups.
- Thirdly, there are then the small matters of character state delimitation and ancestral state reconstruction/character optimisation...
Here there are some general issues to think about first. Sattler (e.g. 1992) has long insisted that our thoughts as it were get trapped inside the terms we use. The most important thing to remember when using botanical terms is that just because some aspect of an organism is dignified by a sesquipedalian word, this by no means signifies that the term refers to an interesting part of "reality" (see also Kirchoff et al. 2008 and references). As Hesse et al. (2009b: p. 27) noted when writing about pollen morphology "Nature itself neither needs categories nor has any knowledge of them" and "categories are artificial and always delimited by an individual or collective convention". Humans make and define botanical terms, and we use them to facilitate communication, although all too often we take them to be "real" - and then they can be as much an impediment to our understanding as anything else. The terms we use, whether in palynology to describe the shape or surface of pollen grains (e.g. Punt et al. 2006; Hesse et al. 2009b), or in general morphology to describe leaf shape (e.g. see Stearn 1992), may well function quite effectively in communication/description while at the same time being inappropriate for use in character optimization - indeed, the caveats in Hesse et al. (2009b), with sections headed "Inherence of Misrepresentation" and "Controversial or Fuzzy Terms", should make one pause for thought. The discussion in E. J. Edwards et al. (2023) about the evolution of CAM and C4 photosynthesis, whether the variation involved is continuous or gappy, and how do we describe evolution here, raises many important issues.
The previous paragraph was by way of a warning... Understanding evolution and phylogenies depends on our associating evolutionary change/novelties with the correct branch of the phylogenetic tree, and these changes/novelties are homologies, unique similarities between structures in different species that reflect their common ancestry; "synapomorphies". The choice of characters and delimitation of their states is a first step to take as we attempt to flesh out phylogenies. For a still useful introduction to the general issue of characters and what they might signify, see the essays in Wagner (2001), and for different ways of thinking about characters, see Kirchoff et al. (2008); Freudenstein (2005) can also be read with profit, while Sattler (1984), Rutishauser and Sattler (1985), Kirchoff et al. (2008), and others outline different ways of thinking about morphology. Here the approach advocated i.a. by Prusinkiewicz and Barbier de Reuille (2010) as they think about plant form - the result of the activity of self-organizing proceses of the plant rather than being under immediate genetic control - is very helpful as an antidote; throw in processes like heterochrony and heterotopy (e.g. Baum & Donoghue 2002), and our appreciation of morphology can change dramatically. Mathews and Kramer (2012) also help us to think about evolution of form and how novelties might develop over the course of evolutionary history. Characters and their states ("anthers" and "introrse"/"extrorse"/"latrorse") as they are generally used come from a careful study of morphology, with similar structures on different organisms being compared. Here Remane's three main criteria of "homology"/similarity - special properties, position, and intermediates - are to be used (see Remane 1952; Kaplan 1997: 1 ch. 1, 2022; c.f. Endress 2011c, p. 122 - "too limited"; Patterson et al. 2023). "Special properties" can include everything from morphology and anatomy to gene expression, and "position" and "special properties" are criteria used when analyzing DNA sequences; for how to relate genotype and phenotype using the concept of persistent homology, a somewhat related issue, see e.g. M. Li et al. (2018 and references) and Larson (2018). However, Remane's criteria may not yield an unambiguous answer as to what a structure "is", even given a solid phylogeny to provide context for the subsequent interpretation of the structure in question. As Endress (2005c) observed, a number of features - position, function, development, shape, anatomy, histology, gene activity, and relationships to other taxa that clearly have petals - can be used to distinguish a petal (for example) from other floral structures. But if a structure has all points of similarity with things that are otherwise called petals bar one, is it thereby not a petal?, or if it has just one, is it a petal? So what about the outer members of the perianth of Nymphaea which may be part green and part coloured on the outside (Warner et al. 2009)?
At the genetic level the situation is equally complex. Genes do not necessarily "do" particular things, rather, gene expression may be a continuous variable. The expression of one gene can be affected by the expression of others, genes are parts of networks, and can be co-opted to act in a variety of different contexts. Thus Maturen et al. (2005) found that floral organ genes (B- and C-class) were expressed in the large, white inflorescence bracts of Cornus (see also Costa et al. 2005), while A-class genes can be expressed in the inflorescence, or even in leaves (Prenner et al. 2011: Euphorbia and Asteraceae). In the inverted repeat loss clade (Fabaceae-Faboideae) FLO/LFY genes, normally floral meristem identity genes, are also expressed in the leaves - which have subtly different morphologies than those of other Fabaceae (Champagne et al. 2007; Wang et al. 2008; Peng et al. 2011; Townsley & Sinha 2012). In Solanum there are similarities at the genetic level between shoot branching and leaf dissection, and development of the abscission zone of the fruit, made up of arrested meristematic cells, is also involved in this regulatory network (Busch et al. 2011; Périlleux et al. 2014). Finally, phenotypes in two organisms may be very similar, but there may have been developmental system drift, the genes controlling the phenotype evolving over time. Are such phenotypes not homologous? There are indeed examples of such situations in animal systems (see McColgan & DiFrisco 2024 for detailed discussion), and although there seem not to be examples yet from plants, this is something to bear in mind.
I have thought a bit about the delimitation of characters and their states (Stevens 2000 for literature), but not nearly enough. As already suggested, one serious problem is that we commonly describe the variation we see, whether in plant architecture, leaf venation, pollen, ovule, nucellus, endosperm, embryo and many other organ systems, as "types" - the solanad type of embryogenesis, Allium type of embryo sac, etc.. Such terms often represent the categorization of simultaneous variation of many features, but the individual features that make them up may be complex, and/or they may vary independently; all this variation effectively gets "lost" when we use these terms. Furthermore, when looking at what we think is a single character, we often find that its states refer to a part of a continuum of variation of that character such as [character] ovule curvature, [states] ovule anatropous vs campylotropous vs ... (and anther opening mentioned above). But worse, we tend to think that evolution is a matter of one "type" of endosperm development (for example) evolving into another. Many of these states have been used in the botanical literature for well over a century, but all too often they are the result of the more or less arbitrary division of a continuum of variation (see Gift & Stevens 1997 for how this works in practice), as with ovule curvature itself. Although such character states may seem to communicate "information", whether they are necessarily suitable for either phylogenetic analysis or understanding evolution (see Linder & Bouchenak-Khelladi 2017) is decidedly questionable. Thus when interpreting the literature (e.g. Stevens 2000, 2006b), and or course this site, too, one should bear such problems in mind. Of course, as character states change, so may our ideas of evolution (e.g. Lamb Frye & Kron 2003; Hibbett 2004), and Tomescu and Groover (2018) provide a nice example of this as they decompose "vascular cambium" into its underlying variables and then think about the evolution of cambium - or, rather, these variables - across all vascular plants.
Furthermore, understanding even current literature can be difficult because the terms that we use often have no fixed definitions - the term "panicle" can be understood only if you know how the author of the publication in which you find the term was using it (see also Endress 2010b). As Rickett (1954: 2, emphasis in original) noted, "To be uncertain whether "glabrous" means "free from hairs and roughness" or only "free from hairs" is as bad as if π [pi] should stand sometimes for the ratio of a circle to its diameter and sometimes for something else; or as if Cu meant sometimes "copper" and sometimes "brass". Yet this is the state of affairs in botany today". Well over a half century later, this is still the state of affairs. In this context, an ontology that is generally accepted by botanists is essential (see also Vogt et al. 2010); the Plant Ontology Consortium is a resource here, but see Kirchoff et al. (2008) for caveats.
The delimitation of states and characters does not necessarily become easier with increasing knowledge. Thus Buzgo et al. (2004) and Matthews and Endress (2005) show how hard it can be to distinguish between e.g. prophylls and other floral structures, while Penet et al. (2005) find that not all monosulcate pollen in monocots has the same developmental pathway, and suggest that therefore such pollen may not have the same ancestral state. There are all intermediates between vessels and tracheids (Schneider & Carlquist 2009; Carlquist & Schneider 2010; Carlquist 2012a), between fully superior and fully inferior ovaries (Soltis & Hufford 2002), floral phyllotaxis may be very labile, especially in magnoliids and members of the ANITA grade (Staedler & Endress 2009), leaves are not always easily distinguishable from other structures (Rutishauser 1999) - and on it goes. But this is in part a level or universe issue. As a comparison increases in extent, what one thought were differences between structures may disappear - and whether this is important, interesting or simply irrelevant will depend on the problem at hand.
For these and other reasons, character states, the 0's and 1's of a morphological data matrix, are all too often some degrees removed from the actual structures observed on individual specimens, furthermore, all too often there is no clear link between these states and specimens/species on which the structures were observed. Specimens are prepared, observations/measurements made, and these somehow become converted to statements about species and then of more inclusive groups (Stevens 1996). Even for this site, it would be have been ideal if observations and literature could have been directly linked at the level of individual species and specimens in a database - this was obvious even as the possibility of APweb was being discussed in the late 1990s. This would allow problems like wrongly assigned taxa to be dealt with easily - remove a taxon from a family, and all information linked to that taxon would be removed at the same time, perfect when dealing with developing phylogenies. Fortunately, this is an area where major changes are underway, changes spurred by the advent of molecular data and what very soon became a mandatory requirement for DNA sequences used in publications to be deposited in the publicly accessible databases like GenBank (founded in 1982) - publication depends on sequences being made available. Resources like the Dryad Digital Repository (it started in 2008) make the various kinds of data that underpin systematic/evolutionary publications accessible. However, as Drew et al. (2013) noted, alignments and trees were very often not deposited, despite encouragements to do so, and this has made earlier studies of less value than they might have been. There has been nothing that has been functioning as a Genbank for morphological data, and it is often impossible to evaluate whether states in a particular study have been delimited satisfactorily; as in much of taxonomy as a whole there are few data in the sense of observations that can be linked to specimens and that can be reanalyzed when necessary. Data matrices can be deposited in places like MorphoBank (O'Leary & Kaufman 2011), and, more generally, in Dryad, and so you can, for example, access the matrix used by Sauquet et al. (2017) in their work on the ancestral angiosperm flower. More extensive data matrices are being developed (e.g. Sauquet 2016: PROTEUS, not yet open access); Sauquet and Magallón (2018) rightly emphasize the importance of a standardized, fully documented, open-access database for plant morphology. Ultimately, fitting basic variation such as measurements, not character states, to a tree allows considerable flexibility in understanding morphology in the context of local phylogenies (see also Stevens 2000; Endress 2005c, but c.f. Weins et al. 2010 for problems here), and this approach has also been used at higher levels, as in a study looking at the evolution of pollen size and shape in Myrtales (Kriebel et al. 2017; see also Parins-Fukuchi 2017).
A good example of the kinds of problems faced are in the description of endosperm morphology and an understanding of its evolution, particularly in monocots. Tobe and Kadokawa (2010) discuss endosperm morphology in terms of cellular and helobial types (Stenar 1925 for early literature; Swamy & Parameswaran 1962a) while Holloway and Friedman (2008) outline possibilities for the evolution of helobial endosperm. However, this distinction, commonly employed in the literature, does not capture the subtleties of endosperm development (see especially Floyd et al. 1999; Floyd & Friedman 2000, 2001 for alternatives; Rudall et al. 2009b). The initial division of the endosperm is highly asymmetric in Araceae, with subsequent cell divisions initially occurring only in the micropylar chamber; the family is considered to have cellular endosperm (Tobe & Kadokawa 2010; see also Maheshwari & Khanna 1957). The chalazal cells here are sometimes massive (e.g. Paremeswaran 1959). Initial asymmetry in endosperm development characterizes helobial endosperm, so Araceae could also be characterized as having a form of helobial endosperm; Acoraceae also show similar asymmetry (Buell 1938). There is also variation in whether the micropylar (e.g. Nymphaeales) or both micropylar and chalazal (Amborella: most from the latter) domains give rise to the bulk of the endosperm. The evolutionary history of nuclear endosperm development is also unclear (Olsen 2004; see e.g. Zou et al. 2001 for subtleties in its development), and there is variation in nuclear endosperm development, as within Poaceae (Leroux et al. 2014). For further discussion about helobial endosperm, see e.g. Kaplan (1981). Shamrov (2021) has recently suggested a classification of endospem variation into types, subtypes and variations, the two types being cellular and helobial, the first division of the endosperm nucleus resulting in two cells (with walls) or just two nuclei respectively... All rather confusing.
The beginning of a solution to such problems is to decompose these "types" into their underlying independent variables. Thus Blackmore et al. (2009) decomposed the pollen "types" used in the description of Asteraceae pollen into no fewer than 52 separate characters, while Dunning et al. (2017) found that an example of a reversal from C4 to C3 photosynthesis did not hold up when the independant variables that make up photosynthetic "types" were analysed (see also Floyd et al. 1999; Floyd & Friedman 2000; Herendeen & Miller 2000; Acosta et al. 2009; Y. Yang et al. 2014; Washburn et al. 2015; et al. 2015, etc.). Character states should be delimited only after analysis of the variation of that character at the level of the study being carried out. Thus delimiting states for a local study in the context of the variation shown by a character across all angiosperms may well be inappropriate since gaps in the variation shown by a particular feature in one clade may be obscured by variation in a completely unrelated clade - the universe problem.
After we have decided on characters and states, there is then the not-so-small question of where character state changes should be placed on the tree, of ancestral state reconstruction/character optimisation. I am fitting characters to conservative trees that may have several polytomies. This makes optimisation of characters, that is, the assigment of character state change to a particular place on the tree, difficult (e.g. Madison & Madison 2002). In nearly all studies of the evolution of characters (D. Soltis et al. 2005b is a good example), the distributions of those characters are optimised on more or less fully resolved trees, and the construction of supertrees may yield more detailed hypotheses of relationships (for supertrees, see e.g. Cotton and Wilkinson 2007, 2008; Buerki et al. 2010d and literature; also see the supermatrix approach). Of course, some nodes even on fully resolved trees and/or supertrees may have little support, and character optimisations will then carry correspondingly little conviction. Topological uncertainties are important here: If Tofieldiaceae are sister to the rest of Alismatales (e.g. Ross et al. 2015), then where the change from "cellular" to "helobial" endosperm should be placed on the tree is unclear. Either one gain (apomorphy for order) and one loss (Araceae), or two gains (Tofieldiaceae, and above Araceae). But the problem is broader, since helobial-type development also occurs in the ANITA grade (e.g. Rudall et al. 2009b). Similarly, is triploid endosperm a synapomorphy for all angiosperms or only for the [[magnoliid + Chloranthaceae] [other angiosperms]] clade (Friedman 2001a, b, 2006; Baroux et al. 2002)? Friedman et al. (2003a, esp. b) and Friedman and Williams (2003, 2004) incline towards the latter hypothesis - see especially Williams and Friedman (2004) and Friedman and Ryerson (2009). If Nymphaeales are sister to Amborellales (but see above), then diploid endosperm may be the ancestral condition, with triploid endosperm evolving in parallel in Amborella and above the ANITA grade - and a supporting argument here might be that Amborella has an embryo sac unique in the angiosperms. Simple parsimony, whether ACCTRAN or DELTRAN, helps little here since there are no outgroups (e.g. Friedman & Floyd 2001; Ronse De Craene et al. 2003). But aside from such uncertainty, even with genomic data trees may remain unresolved in places. In such situations thinking about character evolution is very difficult, putting it mildly (see e.g. Koenen 2019 for the issues involved).
Indeed, using either Bayesian or maximum likelihood analyses, making apparently reasonable suggestions about weighting gains over losses (or vice versa), or more complex assumptions along similar lines, or just using the rather simple models of evolution explicit in ACCTRAN or DELTRAN to place the character on the tree (e.g. Donoghue & Ackerley 1996; Cunningham et al. 1998; Omland 1997, 1999; Ree & Donoghue 1999; Polly 2001; Webster & Purvis 2001; Ronquist 2004; Crisp & Cook 2005; Remizowa et al. 2010b; Sokoloff et al. 2013d; O'Meara 2012; Gascuel & Steel 2014; Wortley et al. 2015; Sauquet et al. 2017; Parkins-Fukuchi 2017) may greatly affect the position of synapomorphies on trees, and hence our ideas of evolution. Sannier et al. (2007) give a good example concerning where on a tree one might peg changes in microsporogenesis in palms (see also Sannier et al. 2009), while Syme and Oakley (2012) note that some methods allow reversals much more easily than others. Pedersen et al. (2007) discuss the sometimes very substantial effect of node support on the posterior probabilities of ancestral character states. But, as Wortley et al. (2015: p. 195) noted rather sadly about optimization, "In general, characters that were unambiguously optimized in all contingencies were either those for which little data were available... or those known to be variable only within a single group, ...". If not reporting on other studies, here I think in terms of parsimony optimization, not always as explicit as it might be, but I have tried to indicate uncertainties as to where particularly important characters change on the tree.
Thus simple parsimony is just one of the ways to optimise the positions of character state changes on a tree, and it may not be the best. Different analyses of the same data may result in state changes ending up in different places on the tree, and/or there may be very different numbers of state changes, and/or changes are interpreted as probabilities given what what we know about the evolution of the character, branch lengths of the tree, topological uncertainties of that tree, and the like. Recent examples are studies of pollen morphology (Wright & Hillis 2014; Wortley et al. 2015; Lu et al. 2015) and of the construction of the ancestral angiosperm flower (Sauquet et al. 2017), where parsimony, maximum likelihood and Bayesian analyses are all compared. Sauquet et al. (2017: Supplementary Methods) capture the issues well, e.g. "MP... has the attractive property that no assumption is made on the relationship between change and branch length; ... Conversely, ML and rjMCMC results are conditional on the assumption that rates of morphological change are constant through times and across lineages." Despite this apparent attractive property of MP, the behaviour of ML and rjMCMC methods was seen as an "advantage" (ibid.). However, molecular clocks are somewhat out of favour these days, and the reasons for invoking a morphological clock, as in ML and rjMCMC methods, are unclear. The bottom line here is that every method has its problems, and extreme care is needed when using them.
A final point is that data must remain easily accessible and should cumulate, that is, ideally information for variation in all characters across all flowering plants (say) should be freely available, be added to as new publications appear, the data being linked to place of publication and specimens examined. In the context of such databases both classifications and the meanings of botanical terms should be standardized. At first sight this might seem an unattainable, almost laughable goal. However, ecologists seem to be showing the way with TRY Plant Trait database (see Kattge et al. 2011; Kissling et al. 2018), traits like wood density, growth form and photosynthetic pathway, which on 20.x.2019 included 11,850,781 trait records from 279,875 species of plants, and more local databases such as PalmTraits 1.0 (Kissling et al. 2019) are being developed - and the data will also be part of TRY. Along these lines, it has long struck me how little the more taxonomic side of our business has to show for over 250 years of work - for instance, most information in the majority of species descriptions cannot be linked to particular specimens. Indeed, thinking of taxaonomy as a cumulative operation, all we have been able to cumulate over the years are names, and we can argue about those until the cows come home. Overstated, of course, but far less so than one would like. However, recent attempts to work out the floral morphology of the ancestral angiosperm (e.g. Sauquet et al. 2017) and to understand various aspects of early angiosperm evolution have involved the construction of data bases that can can be elaborated as work progresses.
The take-home message?: We make the botanical terms we use, not "nature". For me, seeing a plant is forgetting the names its parts are called (with apologies to Weschler 2009); different botanical terms used for what seems to be the same structure are flags that there may be variation out there, but not necessarily much more. As Hesse et al. (2009b: p. 27) observed, "Nature itself neither needs categories nor has any knowledge of categories". More particularly, as Olsen (2004: p. S215) noted, "In spite of recent progress in understanding angiosperm phylogeny, all of the main questions regarding the evolutionary history of the nuclear endosperm remain unresolved." - and this is still the case.
There are six final points to remember about the characterisations below, and descriptions in general:
1. I have been much more generally comparative in the ultimate characterisations than is perhaps strictly necessary. That is, the user will be able to find information on the variation of most characters mentioned in the character list as it occurs in all families. An example is leaf ptyxis, which seems to characterise only a few or the terminal groups recognised here, although I have tried to include information about it more generally.
2. More conventional descriptions of a family, for example, as in Cronquist (1981), include a mixture of plesiomorphies and apomorphies at various levels, including apomorphies of groups of genera, or even individual genera, within the family. Although it is difficult to disentangle the importance of the features listed (but typographic conventions, as in Judd et al. 2002, 2015, help), they better convey a description of the organism as a whole. Nevertheless, by allowing the summarization of the relevant part of the character state hierarchy before each ordinal characterization, I perhaps give something of the same effect.
3. Some of the features in the characterisations simply describe the extent of variation within the clade, e.g., "stomata anomocytic or paracytic". This is particularly common when relationships within a family, or between a family and its immediate relatives, are unclear. Features so described will not often turn out to be synapomorphies, although elements of the variation they encompass may.
4. There may seem to be little "biology" in the lists of characters at the beginning of each page here. In this context, I am reminded of Stebbins's review of Cronquist (1981) which reads in its entirety "The only material of even peripheral interest to the general evolutionist consists of short commentaries on family relationships placed at the end of the description [sic] of many of the families" (Stebbins 1982: p. 628). However, there may be more of interest if you go to the appropriate place in the hierarchy, even if we know little about the functional or adaptive significance of many synapomorphies. Apart from some wind-pollinated, aquatic and parasitic taxa, it is usually difficult to characterise larger groups ecologically, although clades like most of Ericaceae are exceptions. Indeed, much of the "biology" in more conventional descriptions comes from mention of the pollination biology, for example, of particular genera and other small groups within a family, and users can add this emphasis here as they focus on the taxa that grow locally. However, information on more general aspects of ecophysiology, divergence and/or diversification, pollination and fruit dispersal, and associations with particular groups of herbivores, bacteria and fungi, etc., is being added as it becomes available.
5. As we find out more about variation we find fewer and fewer features that are constant throughout a group. Most unqualified statements of presence and absence should properly be qualified as "usually present" or "usually absent" if one is thinking of the characterisations as encompassing the total variation within a clade. Thus Pistia, alone among monocots as so far known, has sieve tube plastids with starch grains, not protein crystals, although the latter is indeed an apomorphy for monocots. Just about all characters that are high-level apomorphies reverse and/or occur in parallel several times on the tree. Thus Reyes et al. (2018) found that monosymmetry originated at least 148 times in angiosperms, and has reversed 69 times or so, and there was widespread parallelism in the other four characters that they examined. Some taxa in Malvaceae-Sterculioideae and Ranunculaceae are effectively gymnospermous...
6. Fossils may have character combinations unknown in extant taxa, as may be seen in the discussions of Fagaceae, Platanaceae, Iteaceae and Calycanthaceae, for example. How this will affect characters at interior nodes remains to be seen. Moreover, where exactly on the tree a particular fossil is to be placed can be difficult to work out - see Nymphaeaceae, Calycanthaceae, Paracryphiaceae, etc., fossils such as Archaefructus and Silvianthemum posing particular problems.
The bottom line is that our morphological, anatomical and chemical knowledge of many critical taxa is woefully incomplete, and much basic - and unfortunately perhaps unfashionable - work must be undertaken to clarify the distribution of morphological, anatomical and chemical characters so we can think more about the evolution of plants. For excellent examples of what can to be done, see the work of P. K. Endress, W. E. Friedman, P. Leins and C. Erbar, L.-P. Ronse de Craene and H. Tobe and all their collaborators (various aspects of floral morphology and development), S. R. Jensen and collaborators (iridoids), and so on, and it is always a pleasure to read papers that change how one thinks about the morphology s.l. of plants (e.g. Matsunaga & Smith 2021: palm fruits). Really remarkable discoveries continue to be made, for instance, the realization that the "typical" 8-ncleate embryo sac is not found in basal angiosperms (e.g. Friedman 2001a - for "basal", see below), and the description of the embryology of Cardiopteris, related to holly, that shows it to be without parallel in any other flowering plant (Tobe 2016). Indeed, filling in the gaps in our knowledge of more mundane things like nodal anatomy, cell and tissue distributions, stomatal morphology, seed coat anatomy, and the like can be quite easy (see also below) and will clarify our understanding of evolution. Much effort must continue to be spent in summarizing characters of well-established clades at all levels, providing features by which they may be recognized, and signaling synapomorphies. However, given our current understanding of both phylogenies and morphology, how to relate the two is not that simple, and different assumptions about character evolution may greatly affect the position of synapomorphies on trees. Nevertheless, it can be a relatively easy matter to update notes such as these and to incorporate data on characters that have never before been considered in the context of a phylogenetic ttree - even one in which there has been grafting between the branches.
SUMMARY OF APG IV, MAIN CHANGES SINCE, LINKS TO PAGES.
[Back to Top]
Below is a summary of the relationships within orders of all the families of seed plants; there are direct links to the pages where the families are discussed. Families, etc., accepted in A.P.G. IV (2016) and the most recent edition of The Plant Book (Mabberley 2017) are very largely the same as those below.
However, there are a few changes in relationships and new families/orders that have been made since APG IV. The latter are listed immediately below, while in the summary they are indicated by asterisks. Some of these changes are new families that reflect recent collections, e.g. the remarkable Tiganophytaceae, from the Namib area and in Brassicales, while others, like Oncothecales, a very small order that may well be sister to all other core asterids/gentianids, are the results of changed ideas of relationships that are being suggested by comprehensive analyses using nuclear genomes. This latter kind of change in particular is being made only when support for the changed relationships is strong, otherwise confusion will result. However, now that PAFTOL is off the ground, with an immediate goal of including all genera of flowering plants (W. J. Baker et al. 2021a, b: see also Seed Plant Tree of Life - most reecent version Jan. 2022), who knows what the future holds? Note that many relationships along the spine of the Pentapetalae well up into the asterids and within orders like Malpighiales are subject to change. Those changes that are being made are being carefully justified.
Square brackets - [...] - enclose clades; the plus sign - + - designates terminal sister taxa; a comma - , - denotes part of a polytomy.
MAJOR ADDITIONS/CHANGES SINCE 2016
Brassicales: add Tiganophtyaceae.
Santalales: add Mystropetalaceae.
Core Asterids/gentianids: add Oncothecales: Oncothecaceae.
Lamiales: add Wightiaceae
CAMPANULID/ASTERID II: Add Cardiopteridales: [Cardiopteridaceae + Stemonuraceae]
Add: Desfontainiales: Columelliaceae
Note also that the circumscriptions of Aquifoliales, Bruniales, Icacinales and Metteniusales have changed, relationships along the spine of the tree above Gunnerales and the Ericornids, and also in Ericales and Malpighiales (just to name two prominent orders) are up for grabs, etc..
SEED PLANTS
GYMNOSPERMS
Cycadales
[Cycadaceae + Zamiaceae]
Ginkgoales[[Araucariaceae + Podocarpaceae] [Sciadopityaceae [Taxaceae + Cupressaceae]]]
Pinales[Ephedraceae [Gnetaceae + Welwitschiaceae]]
ANGIOSPERMS/FLOWERING PLANTSEvolution and diversification of the angiosperms
Amborellales[Hydatellaceae [Cabombaceae + Nymphaeaceae]]
Austrobaileyales[Austrobaileyaceae [Trimeniaceae + Schisandraceae]]
Mesangiosperms[Myristicaceae [Magnoliaceae [[Himantandraceae + Degeneriaceae] [Eupomatiaceae + Annonaceae]]]]
Laurales[Calycanthaceae [[Siparunaceae [Gomortegaceae + Atherospermataceae]] [Lauraceae [Monimiaceae + Hernandiaceae]]]]
Canellales[Aristolochiaceae [Piperaceae + Saururaceae]]
[Araceae [Tofieldiaceae [[Alismataceae [Hydrocharitaceae + Butomaceae]] [Scheuchzeriaceae [Aponogetonaceae [Juncaginaceae [Maundiaceae [[Posidoniaceae [Ruppiaceae + Cymodoceaceae]] [Zosteraceae + Potamogetonaceae]]]]]]]]]
PetrosavialesPetrosaviaceae [?Here. ?Asparagales]
Dioscoreales[Nartheciaceae [[Taccaceae + Thismiaceae (limits?)] [Burmanniaceae + Dioscoreaceae]]]
Pandanales[Velloziaceae [Stemonaceae [Triuridaceae [Pandanaceae + Cyclanthaceae]]]]
Liliales[[Corsiaceae + Campynemataceae] [[Petermanniaceae [Colchicaceae + Alstroemeriaceae]] [Melanthiaceae [[Liliaceae + Smilacaceae] [Philesiaceae + Ripogonaceae]]]]]
Asparagales[Orchidaceae [[Boryaceae [Blandfordiaceae [Asteliaceae [Lanariaceae + Hypoxidaceae]]]] [[Ixioliriaceae + Tecophilaeaceae] [Doryanthaceae [Iridaceae [Xeronemataceae [Asphodelaceae [Amaryllidaceae + Asparagaceae]]]]]]]]
COMMELINIDS[[Typhaceae + Bromeliaceae] [Rapateaceae[Mayacaceae [Eriocaulaceae + Xyridaceae] [Thurniaceae [Juncaceae + Cyperaceae]] [Restionaceae [Flagellariaceae [Joinvilleaceae [Ecdeiocoleaceae + Poaceae]]]]]]]]
Commelinales[[Philydraceae [Commelinaceae + Hanguanaceae]] [Haemodoraceae + Pontederiaceae]]
Zingiberales[[Musaceae [Heliconiaceae [Strelitziaceae + Lowiaceae]]] [[Cannaceae + Marantaceae] [Costaceae +Zingiberaceae]]]
[Eupteleaceae [Papaveraceae [[Lardizabalaceae + Circaeasteraceae] [Menispermaceae [Berberidaceae + Ranunculaceae]]]]]
Proteales[Sabiaceae [Nelumbonaceae [Platanaceae + Proteaceae]]]
Trochodendrales[Gunneraceae + Myrothamnaceae]
PENTAPETALAE[Peridiscaceae [[Paeoniaceae [Altingiaceae [Hamamelidaceae [Cercidiphyllaceae + Daphniphyllaceae]]]] [[Crassulaceae [Aphanopetalaceae [Tetracarpaeaceae [Penthoraceae + Haloragaceae]]]] [Iteaceae [Grossulariaceae + Saxifragaceae]]]]], Cynomoriaceae unplaced
ROSIDS[Krameriaceae + Zygophyllaceae]
Oxalidales[Huaceae [[Connaraceae + Oxalidaceae] [Cunoniaceae [Cephalotaceae [Elaeocarpaceae + Brunelliaceae]]]]]
Celastrales[Lepidobotryaceae + Celastraceae]
Malpighiales[[Ctenolophonaceae [Erythroxylaceae + Rhizophoraceae]], Irvingiaceae, Pandaceae, [Ochnaceae [[Bonnetiaceae + Clusiaceae] [Calophyllaceae [Hypericaceae + Podostemaceae]]]]] [[[Lophopyxidaceae, Putranjivaceae], Caryocaraceae, Centroplacaceae, [Elatinaceae + Malpighiaceae], [Balanopaceae [[Trigoniaceae + Dichapetalaceae] [Chrysobalanaceae + Euphroniaceae]]]] [[Humiriaceae [[Achariaceae [Goupiaceae + Violaceae] [Passifloraceae [Lacistemataceae + Salicaceae]]]]] [[Peraceae [Rafflesiaceae + Euphorbiaceae]] [[Phyllanthaceae + Picrodendraceae] [Linaceae + Ixonanthaceae]]]]]
N-FIXING CLADE[Quillajaceae [Fabaceae [Polygalaceae + Surianaceae]]]
Rosales[Rosaceae [[Rhamnaceae [Elaeagnaceae [Barbeyaceae + Dirachmaceae]]] [Ulmaceae [Cannabaceae [Moraceae + Urticaceae]]]]]
Cucurbitales[Anisophylleaceae [[Corynocarpaceae + Coriariaceae] [Cucurbitaceae [Tetramelaceae [Datiscaceae + Begoniaceae]]]], Apodanthaceae unplaced
Fagales[Nothofagaceae [Fagaceae [[Myricaceae + Juglandaceae] [Casuarinaceae [Ticodendraceae + Betulaceae]]]]]
MALVIDS/ROSID IICombretaceae, [Onagraceae + Lythraceae], [[Vochysiaceae + Myrtaceae] [Melastomataceae [Crypteroniaceae [Alzateaceae + Penaeaceae]]]]
Crossosomatales[[Staphyleaceae [Guamatelaceae [Crossosomataceae + Stachyuraceae]]] [Aphloiaceae [Geissolomataceae + Strasburgeriaceae]]]
Picramniales[Biebersteiniaceae, Nitrariaceae, [[Kirkiaceae [Anacardiaceae + Burseraceae]] [Sapindaceae [Meliaceae [Simaroubaceae + Rutaceae]]]]]
Huerteales[[Gerrardinaceae + Petenaeaceae] [Tapisciaceae + Dipentodontaceae]]
Malvales[Neuradaceae [Thymelaeaceae [Sphaerosepalaceae, Bixaceae, [Cistaceae [Sarcolaenaceae + Dipterocarpaceae]], [Cytinaceae + Muntingiaceae], Malvaceae]]]
Brassicales[[Akaniaceae + Tropaeolaceae] [[Moringaceae + Caricaceae] Setchellanthaceae[ [Limnanthaceae [[Koeberliniaceae, Bataceae, Tiganophtyaceae, Salvadoraceae] [Emblingiaceae [[Pentadiplandraceae [Gyrostemonaceae + Resedaceae]], Tovariaceae, [Capparaceae [Cleomaceae + Brassicaceae]]]]]]]]]]
[Erythropalaceae [Strombosiaceae [Coulaceae [[Ximeniaceae [Aptandraceae + Olacaceae]] [Octoknemaceae [Balanophoraceae [[[Mystropetalaceae + Loranthaceae] [Misodendraceae + Schoepfiaceae]] [Opiliaceae + Santalaceae]]]]]]]]]
Berberidopsidales[Aextoxicaceae + Berberidopsidaceae]
Caryophyllales[[[Droseraceae + Nepenthaceae] [Drosophyllaceae [Ancistrocladaceae + Dioncophyllaceae]]] [[Frankeniaceae + Tamaricaceae] [Polygonaceae + Plumbaginaceae]]] [Rhabdodendraceae [Simmondsiaceae [[Asteropeiaceae + Physenaceae] [Macarthuriaceae [Microteaceae [[Caryophyllaceae [Achatocarpaceae + Amaranthaceae]] [Stegnospermataceae [Limeaceae [[Lophiocarpaceae [Kewaceae [Barbeuiaceae [Aizoaceae [Gisekiaceae [[Sarcobataceae + Phytolaccaceae] [Petiveriaceae + Nyctaginaceae]]]]]]] [Molluginaceae [Montiaceae [[Halophytaceae [Didiereaceae + Basellaceae]] [Talinaceae [Portulacaceae [Anacampserotaceae + Cactaceae]]]]]]]]]]]]]]]]
ASTERIDS[[Cornaceae [Grubbiaceae + Curtisiaceae]] [Nyssaceae [Hydrostachyaceae [Hydrangeaceae + Loasaceae]]]]
Ericales[[Balsaminaceae [Marcgraviaceae + Tetrameristaceae]] [[Polemoniaceae + Fouquieriaceae], Lecythidaceae, [[Sladeniaceae + Pentaphylacaceae], [Sapotaceae [Ebenaceae + Primulaceae]], [Mitrastemonaceae, Theaceae, [Symplocaceae [Styracaceae + Diapensiaceae]], [[Sarraceniaceae [Roridulaceae + Actinidiaceae]] [Clethraceae [Cyrillaceae + Ericaceae]]]]]]]
EUASTERIDS[Aquifoliaceae [Helwingiaceae + Phyllonomaceae]]
Icacinales[Rubiaceae [[Loganiaceae + Gelsemiaceae] [Gentianaceae + Apocynaceae]]]
Vahliales[[Montiniaceae [Sphenocleaceae + Hydroleaceae]] [Convolvulaceae + Solanaceae]]
Boraginales[[Codonaceae [Wellstediaceae + Boraginaceae]] [Hydrophyllaceae [Namaceae [Heliotropiaceae [Cordiaceae + Ehretiaceae]]]]
Lamiales[Plocospermataceae [[Carlemanniaceae + Oleaceae] [Tetrachondraceae [[Peltantheraceae [Calceolariaceae + Gesneriaceae]] [Plantaginaceae [Scrophulariaceae [Stilbaceae [[Byblidaceae + Linderniaceae] [[Thomandersiaceae + Verbenaceae], Pedaliaceae, Schlegeliaceae, Martyniaceae, Bignoniaceae, Acanthaceae, Lentibulariaceae] [Lamiaceae [Mazaceae [Phrymaceae [[Paulowniaceae + Wightiaceae] + Orobanchaceae]]]]]]]]]]]]]
CAMPANULID/ASTERID II[Cardiopteridaceae + Stemonuraceae]
Escalloniales[[Rousseaceae + Campanulaceae] [Pentaphragmataceae [Stylidiaceae [[Alseuosmiaceae [Phellinaceae + Argophyllaceae]] [Menyanthaceae [Goodeniaceae [Calyceraceae + Asteraceae]]]]]]]
Bruniales[Pennantiaceae [Torricelliaceae [Griseliniaceae [Pittosporaceae [Araliaceae [Myodocarpaceae + Apiaceae]]]]]]
Paracryphiales[Viburnaceae + Caprifoliaceae]
ON SOME POORLY-KNOWN TAXA THAT ARE IN URGENT NEED OF STUDY
[Back to Top]
Although a vast amount of morphological s.l. information is being developed from question-driven research on model organisms like Zea, Arabidopsis and Antirrhinum, it is often difficult to know how far the results can be extrapolated. Indeed, I have long been impressed (or depressed) about several aspects about our general knowledge (including my own) of morphology.
— How little we really know when we discuss evolution - for most features, we extrapolate from a quite small data base.
— How little morphological s.l. knowledge is added in most phylogenetic studies.
— How much we are dependent on general surveys from the 1880s to the 1930s in particular (for cytology and plant chemistry the surveys are later, but even here our knowledge base is increasing very slowly), although for aspects of floral morphology and development, for example, a few authors have added substantial amounts of data over the last thirty years. Fortunately, much of this older literature is now on line, for example, see the Botanicus Digital Library, with approaching 2,500,000 pages scanned, while in the Biodiversity Heritage Library there are about 46,000,000 pages [as of 13.iv.2015].
— How easy it is to add new information about many organ systems.
A more comprehensive understanding of plant morphology is needed to enable broad evolutionary questions to be answered with a greater degree of confidence. There is a particular need for targeted surveys of the morphological variation of particular groups. Given limited time and resources, one can also profitably focus on small clades an understanding of the morphology of which will have the greatest effect on our understanding of character evolution. These include taxa along the spines of branches, a small clade sister to the rest of a very speciose clade, etc.. Add to that a judicious sampling of more speciose clades (see also Endress 2011) carefully targeted in the context of the phylogeny of the group of interest; untargeted surveys, or surveys of groups whose phylogeny is unclear, may yield less of interest than one might hope (c. f. Schönenberger & von Balthazar 2006). Neither phylogeny without good comparative morphological knowledge nor comparative morphology without a good phylogeny is of much use.
To help in building up a more extensive information base that will facilitate our thinking about the larger patterns of flowering plant evolution, the list below includes a number of quite small taxa whose phylogenetic position is such that information on embryo sac development, chemistry, seed anatomy, etc., is particularly important for our understanding of the evolution of those features. The taxa listed are reasonable targets for comprehensive studies and are eminently suitable for Masters theses. Of course, given the current academic climate, I cannot say that such studies will be highly regarded by tenure and promotion committees (bless their hearts, wherever they might be), but they should nevertheless be integral to all phylogenetic studies that are concerned with character evolution and integral parts of grant proposals.
The taxa listed below come from all over the world except the northern Temperate and Arctic zones. Tropical Africa (even excluding the Cape region and Madagascar), Central and tropical South America, and Australia-New Zealand are particularly well represented, East Asia and Indo-Malesia somewhat less so, then the Cape region, Madagascar, and the southern U.S.A./northern Mexico, and finally New Caledonia, temperate South America, and Europe. Although some of these genera like Triceratella (Commelinaceae) are very rare, others are quite common, or are even in cultivation (Rehmannia, Aextoxicon, Portulaca, etc.). Such a list does not pretend to be exhaustive, indeed, there are some genera that are quite well known, but for which important information is lacking - thus I know of no reports of fertilization from Nothofagus (Nothofagaceae, sister to rest of Fagales). Given our general level of knowledge, there is no shortage of plants to look at!
In addition to the taxa listed, many others, especially in Geraniales, practically the whole of Phytolaccaceae and Molluginaceae, Irvingiaceae, Kirkiaceae, Acanthaceae-Nelsonioideae, Escalloniaceae, Bruniaceae, etc., could well have been included, and as phylogenies in Lamiales, etc., become better resolved, other clades in urgent need of study will become apparent. Many holoparasites need developmental and embryological studies.
Mention of “basal” in the comments below is relative; it means basal in a tree relative to other more speciose clades or simply to other clades that are being discussed - trees, even phylogenetic trees, have bases. There is no implication that members of such basal clades are “primitive” or plesiomorphic (c.f. McDaniel 2021).
Achatocarpaceae: Caryophyllales. Achatocarpus (Mexico to Argentina) Phaulothamnus (S.W. U.S.A., N. Mexico): 10 and 1 spp. respectively. Sister to Amaranthaceae s.l.
Aextoxicaceae: Berberidopsidales. Aextoxicon (Chile): 1 sp. One of two families in Berberidopsidales, along spine below the asterids, see also Berberidopsidaceae.
Aizoaceae: Caryophyllales. Acrosanthes (Western Cape, South Africa): 6 spp. The only genus in its subfamily.
Akaniaceae: Brassicales. Akania (E. Australia), Bretschneidera (S.E. Asia): 1 and 1 spp. respectively. With Tropaeolaceae, sister to all other Brassicales.
Alseuosmiaceae: Asterales. Alseuosmia (New Zealand), Crispiloba (N.E. Australia), Periomphale (New Caledonia), Platyspermation (New Caledonia), Wittsteinia (S.E. Australia, New Guinea): 5, 1, 1, 1, and 2 spp. respectively.
Anacampserotaceae: Caryophyllales. Anacampseros, Avonia, Grahamia (Africa, S.W. U.S.A., Central and South America, Australia, very scattered): 32 spp. in whole family. Sister to Cactaceae.
Aphanopetalaceae: Saxifragales. Aphanopetalum (W. and E. Australia): 2 spp.
Aphloiaceae: Crossosomatales. Aphloia (E. Africa, Madagascar, Seychelles): 1-several spp. Sister to one major clade in the order.
Asteraceae: Asterales. Corymbium (South Africa); 7 spp. Sister to Asteroideae.
Asteropeiaceae: Caryophyllales. Asteropeia (Madagascar): 8 spp. One family of two in clade sister to core Caryophyllales, see also Physenaceae.
Barbeuiaceae: Caryophyllales. Barbeiua (Madagascar): 1 sp. Sister to a very speciose part of Caryophyllales.
Barbeyaceae: Rosales. Barbeya (Horn of Africa, Arabia): 1 sp. Near Rhamnaceae.
Berberidopsidaceae: Berberidopsidales. Streptothamnus (E. Australia): 1 sp., florally very different from Berberidopsis. One of two families in Berberidopsidales, along spine below the asterids, see also Aextoxicaceae.
Bixaceae: Malvales. Diegodendron (Madagascar): 1 sp. Isolated in Bixaceae.
Brassicaceae: Brassicales. Aethionema (Mediterranean to Afghanistan): 70 spp. Sister to all other Brassicaceae.
Buxaceae: Buxales. Didmyeles (Madagascar): 2 spp. Sister to rest of family.
Carlemanniaceae: Lamiales - sister to Oleaceae, together sister to the rest of the order, minus one species! Carlemannia, Silvianthus (both S.E. Asia to W. Malesia): 3 and 1 spp. respectively.
Campanulaceae: Asterales. Cyphioideae: Cyphia (Africa): 60 spp. Cyphocarpoideae: Cyphocarpus (Chile): 2 spp. Nemacladoideae: Nemocladus (S.W. North America): 12 spp. Basal or nearly so in the family.
Campynemataceae: Liliales. Campynema (Tasmania), Campynemanthe (New Caledonia, somewhat known): 1 and 3 spp. respectively.
Caprifoliaceae: Dipsacales. Zabelia (East Asia, Afghanistan to Japan): 6 spp. Crucial position in middle of Caprifoliaceae.
Caryocaraceae: Malpighiales. Anthodiscus (tropical South America), Caryocar (Central America, tropical South America): 8 and 15 spp. respectively.
Centroplacaceae: Malpighiales. Centroplacus (West Africa), Bhesa (Indomalesia): 1 and 5 spp. respectively.
Codonaceae: Boraginales. Codon (South Africa): 2 spp. Sister to clade including Boraginaceae.
Combretaceae: Myrtales. Strephonema (West Africa): 3 spp. Sister to rest of family.
Commelinaceae: Commelinales. Cartonema (N. and S.W. Australia, New Guinea) and probably Triceratella (Zimbabwe): 11 and 1 spp. respectively. In clade sister to rest of family.
Convolvulaceae: Solanales. Humbertia (Madagascar): 1 sp. Sister to rest of family.
Cordiaceae: Boraginales. Hoplestigma (West Africa). 1 spp. Strange floral morphology.
Ctenolophonaceaee: Malpighiales. Ctenolophon (tropical West Africa, Malesia): 3 spp.
Cucurbitaceae: Cucurbitales. Indofevillea (N. E. India, Assam, Tibet, Myanmar): 2 spp. Sister to all other Cucurbitoideae.
Curtisiaceae: Cornales. Curtisia (Southern Africa): 1 sp.
Cyrillaceae: Ericales. Cliftonia (S.E. U.S.A), Cyrilla (S. U.S.A. to N. South America): 1 and 1 spp. respectively. Sister to Ericaceae.
Dasypogonaceae: Arecales. Baxteria (S.W. Australia), Calectasia (S.E. Australia), Dasypogon (S.W. Australia), Kingia (S.W. Australia): 1, 11, 3, and 1 spp. respectively. ?Sister to Arecaceae.
Dipentodontaceae: Huerteales. Dipentodon (E. Asia), Perrottetia (Central and W. South America, S.E. Asia to Australia): 1 and 15 spp. respectively. All Huerteales poorly known.
Dirachmaceae: Rosales. Dirachma (Horn of Africa): 2 spp. Near Rhamnaceae.
Euphroniaceae: Malpighiales. Euphronia (N. South America): 3 spp.
Fabaceae: Fabales. Duparquetia (tropical West Africa): 1 sp. Near basal in family.
Geissolomataceae: Crossosomatales. Geissoloma (South Africa): 1 sp.
Gelsemiaceae: Gentianales. Pteleocarpa (W. Malesia): 1 sp. Distinctly odd.
Geraniaceae: Geraniales. Hypseocharis (Andean South America): 1-3 spp. Sister to rest of family.
Gerrardinaceae: Huerteales. Gerrardina (S. and E. Africa): 2 spp.; they look rather different. All Huerteales are poorly known.
Giseckiaceae: Caryophyllales. Giseckia (Africa to E. Asia): 5 spp. Core Caryophyllales.
Goupiaceae: Malpighiales. Goupia (Central and N.E. South America): 2 spp. Probably sister to Violaceae.
Griseliniaceae: Apiales. Griselinia (New Zealand and S. South America): 6 spp.
Grubbiaceae: Cornales. Grubbia (South Africa, the Cape): 3 sp.
Guamatelaceae: Crossosomatales. Guamatela (Guatemala): 1 sp.
Huaceae: Oxalidales. Afrostyrax, Hua (both tropical W. Africa): 1 and 2 spp. respectively. Perhaps sister all other Oxalidales.
Icacinaceae: Icacinales. Includes Casimirella (Paraguay, 1 sp.), Cassinopsis (Africa, Madagascar, 6 spp.), Hosiea (China, Japan, 2 spp.), Mappia (tropical America, 5 spp.), Merrillodendron (Philippines, etc., 1 sp.), Nothapodytes (East Asia to Malesia, 4 spp.), Sarcostigma (S.E. Asia, W. Malesia, 2 spp.). Basically the whole family...
Kewaceae: Caryophyllales. Kewa (ex Hypertelis) (Africa, Saint Helena): 8 spp. Near Aizoaceae, core Caryophyllales.
Lacistemataceae: Malpighiales. Lacistema, Lozania (both Central and South America): 11 and 3 spp. respectively. Sister to Salicaceae.
Leycthidaceae: Ericales. Foetidia (centred on Madagascar): 17 spp. Napoleonoideae, sister to rest of family - Crateranthus, Napoleona (both tropical West Africa): 3 and 8 spp. respectively.
Lepidobotryaceae: Celastrales. Lepidobotrys (tropical W. Africa), Ruptiliocarpon (C. America, W. South America): 1 and 1 spp. respectively. Sister to Celastraceae s.l.
Limeaceae: Caryophyllales. Limeum (Africa to Pakistan, esp. South Africa), Macarthuria (S.E./S.W. Australia): 26 and 10 spp. respectively. Clade along spine of core Caryophyllales.
Lophiocarpaceae: Caryophyllales. Lophiocarpus (South Africa), Corbichonia (Africa to Asia): 4 and 2 spp. respectively. Sister to speciose clade in core Caryophyllales.
Lophopyxidaceae: Malpighiales. Lophopyxis (Malesia to W. Pacific): 1 sp.
Mayacaceae: Poales. Mayaca (West Africa, tropical America): 4 spp.
Metteniusaceae: Metteniusales. Includes Apodytes (Old World tropics, 15 spp.), Calatola (Mexico to Ecuador, 7 spp.), Dendrobangia (tropical South America, 2 spp.), Emmotum (tropical South America, 12 spp.), Metteniusa (Costa Rica to NW South America, 7 spp.), Platea (Malesia, 5 spp.), Rhaphiostylis (tropical W. Africa, 6 spp.). Again, basically the whole family.
Microteaceae: Caryophyllales. Microtea (Central and South America, Antilles): 9 spp. Perhaps sister to all other core Caryophyllales minus Macarthuriaceae.
Montiniaceae: Solanales. Grevea (E. Africa, Madagascar), Kaliphora (Madagascar), Montinia (S. Africa): 3, 1, and 1 spp. respectively.
Muntingiaceae: Malvales. Dicraspidia (Central to N.W. South America), Muntingia (tropical America, somewhat known): 1 and 1 sp. respectively. Neotessmannia (Peru): 1 sp. Does this belong?
Neuradaceae: Malvales. Grielium (South Africa), Neurada (E. Mediterranean to India), Neuradopsis (S.W. Africa): 5, 1 and 3 spp. respectively. ?Sister to all other Malvales.
Nitrariaceae: Sapindales. Nitraria (Mediterranean to C. Asia, S. North America), Peganum, Tetradiclis (both E. Europe to C. Asia): 12, 6 and 1 spp. respectively.
Nyctaginaceae: Caryophyllales. Caribea litoralis (Cuba). Sister to rest of family.
Oncothecaceae: Icacinales. Oncotheca (New Caledonia): 2 spp. Sister to all other core asterids.
Orobanchaceae: Lamiales. Rehmannia (East Asia), Trianeophora (EastAsia): 6 and 3 spp. respectively. Sister to rest of family.
Passifloraceae: Malpighiales. Pibiria (Guyana): 1 sp. Sister to Turneroideae.
Paracryphiaceae: Paracryphiales. Paracryphia (New Caledonia), Quintinia (Philippines to New Zealand), Sphenostemon (Celebes to N.E. Australia and New Caledonia); 1, 25, and 10 spp. respectively. Sister to Apiales.
Pennantiaceae: Apiales. Pennantia (E. Australia to New Zealand): 4 spp. Sister to all other Apiales.
Pentadiplandraceae: Brassicales. Pentadiplandra (tropical W. Africa): 1 sp. Core Brassicales.
Pentaphragmataceae: Asterales. Pentaphragma (S.E. Asia-Malesia): 30 spp. Sister to major clade in order.
Peridiscaceae: Saxifragales. Medusandra, Soyauxia (both tropical W. Africa), Peridiscus (Amazonia), Whittonia (N. South America): 2, 7, 1, and 1 spp. respectively. Sister to all other Saxifragales.
Petenaeaceae: Huerteales. Petenaea (C. America): 1 sp. Order poorly known.
Phellinaceae: Asterales. Phelline (New Caledonia): 12 spp.
Phyllonomaceae: Aquifoliales. Phyllonoma (Mexico to W. South America): 4 spp.
Picramniaceae: Picramniales. Alvaradoa (Florida, Central America, Bahama, esp. the Greater Antilles, Bolivia to Argentina), Nothotalisia (Panama and N.W. South America), Picramnia (Florida, Central and South America): 5, 3, and 41 spp. respectively. Along spine of malvids.
Poaceae: Poales. Pharoideae: Leptaspis (Old World tropics), Pharus (New World, Florida to Argentina): 4 and 8 spp. respectively. Puelioideae: Guaduella, Puelia (both tropical W. Africa): 6 and 5 spp. respectively. Clades along basal spine of family.
Physenaceae: Caryophyllales. Physena (Madagascar): 2 spp. One family of two in clade sister to core Caryophyllales, see also Asteropeiaceae.
Plocospermataceae: Lamiales. Plocospermum (Central America): 1 sp. Sister to all other Lamiales.
Picrodendraceae: Malpighiales. Podocalyx (Amazonian South America): 1 sp. Sister to rest of family.
Portulacaceae: Caryophyllales. Portulaca (pan(sub)tropical): 40-100 spp. Once-removed from sister to Cactaceae.
Primulaceae: Ericales. Maesa (Old World Tropics): 160 spp. Sister to rest of family.
Quillajaceae: Fabales. Quillaja (temperate South America): ca 3 spp.
Resedaceae: Forchhammeria (S.W. North America), Stixis (E. Himalayas to W. Malesia), Neothorelia (Laos), Tirania (Vietnam): 10, 7, 1 and 1 spp. respectively. Also Borthwickia (S.W. Yunnan, China, and adjacent Burma): 1 sp.
Rhizophoraceae: Malpighiales. Paradrypetes (tropical South America): 2 spp. Very odd genus.
Rousseaceae: Asterales. One clade includes Roussea (Mauritius): 1 sp. The other clade includes Abrophyllum (E. Australia), Carpodetus (New Guinea to New Zealand), Cuttsia (E. Australia): 1, 2, and 1 spp. respectively. Sister to Campanulaceae.
Sapindaceae: Sapindales. Xanthoceras (China): 1 sp. Possibly sister to rest of family
Sarcobataceae: Caryophyllales. Sarcobatus (S.W. North America): 2 spp. Core Caryophyllales, position unclear.
Setchellanthaceae: Brassicales. Setchellanthus (S.W. U.S.A., Mexico): 1 sp. Along spine of Brassicales.
Simmondsiaceae: Caryophyllales. Simmondsia (S.W. U.S.A., Mexico): 1 sp. Along spine just below core Caryophyllales.
Sladeniaceae: Ericales. Ficalhoa (East Africa), Sladenia (S. E. Asia): 1 and ?2 spp. respectively.
Stegnospermataceae: Caryophyllales. Stegnospermum (Central America, Antilles): 3 spp. Along spine of core Caryophyllales.
Surianaceae: Fabales. Cadelia (N.E. Australia), Guilfoylia (E. Australia), Recchia (Mexico), Stylobasium (W. and N. Australia), Suriana (pantropical, a little known): 1, 1, 3, 2 and 1 spp. respectively. Position in order unclear.
Tapisciaceae: Huerteales. Huertea (Central and South America), Tapiscia (E. Asia): 4 and 1 spp. respectively. Whole order poorly known
Tetracarpaeaceae: Saxifragales. Tetracarpaea (Tasmania): 1 sp.
Tetrachondraceae: Lamiales. Polypremum (New World), Tetrachondra (New Zealand, Chile): 1 and 2 spp. respectively. Sister to very speciose monosymmetric Core Lamiales.
Ticodendraceae: Fagales. Ticodendron (Central America): 1 sp.
Torricelliaceae: Apiales. Aralidium (W. Malesia), Melanophylla (Madagascar), Torricellia (Southeast Asia): 1, 7, and 2 spp. respectively. Sister to all Apiales minus Pennantiaceae.
Vahliaceae: unplaced asterid I. Vahlia (Africa, Madagascar, Indian subcontinent): 8 spp.
Wellstediaceae: Boraginales. Wellstedia (N.E. Africa, South West Africa): 6 spp. Sister to Boraginaceae.
Xeronemataceae: Asparagales. Xeronema (New Zealand, New Caledonia): 1 sp. Along spine of Asparagales.
ON THE ORGANIZATION AND DESIGN OF THE SITE
[Back to Top]
This website is best viewed using the most recent version of your preferred internet browser. You can tell when each page was last updated if you look at the top of each page, or, for the individual order pages, immediately below mention of the order or other major clade first mentioned on that page - thus on the Arecales page the update indication is below the characterization of the commelinid group.
The left pane is designed as a quick entry into different parts of the seed plant phylogeny. The default pane includes a reference list of accepted orders as well as a listing of abbreviations, etc., used in the body of the text. There are also direct links to the developing essays on seed plant evolution and angiosperm evolution and diversification. This left pane can be replaced by a list of the main characters that are described in detail elsewhere - see the "Characters" option in the top pane of the website.
The top pane is designed as a menu:
- Home will always return the user to this page.
- Tree gets you to the Main Tree that displays a phylogeny of the orders. Clicking once on the name of a terminal taxon will take you to a characterization of that taxon. Clicking on a node will take you to that particular node. Clicking on one of the tree icons next to an ordinal name will usually take you a tree showing relationships within that order. Not all orders have such icons; some are too small. Clicking on "Unplaced Taxa" will send the user to those few families and genera that lack any convincing evidence as to their likely immediate relationships.
- Orders provides an alphabetical listing of all accepted ordinal names and also all synonyms. Clicking on a letter at the top of the page will take you to the beginning of names that start with that particular letter.
- Families is an alphabetical listing of all accepted family names and their synonyms. Clicking on a letter at the top of the page will take you to the beginning of names that start with that particular letter. Clicking on a link to a family will take you to its characterization.
- Characters provides a textual description of characters used to support the recognition and/or monophyly of a particular taxon. General references that are important sources of data for this project are also included. At the beginning of this page is a notice "click here"; this will display a list of the major characters included in the left pane. One can return to the Orders list by choosing the option "Back to Orders" at the beginning of the character list.
- References provide an alphabetical listing of the references cited; it has been broken down into six interlinked pages for ease of loading. Clicking on a letter at the top or the bottom of the page will take you to the part of the reference list where surnames with that letter begin. Articles with two authors are listed alphabetically by name of second author and then by date, articles with three or more authors are cited last (e.g. as "Chase [et al. 1993]") and are listed by date. You can also search on individual authors' names.
- "Search" takes you to a page where you may enter keywords or terms that you are interested in finding within the Angiosperm Phylogeny Website. The search engine is unfortunately not designed to search for wildcards, so if your search is not successful, please try variations of your search terms. [Para. to be changed.]
- Links lists some other websites or web utilities that are relevant to general classification, floras (a few), image resources, and phylogeny. These listings are very incomplete. There are also some links to other websites in the Bibliography and on individual ordinal pages.
- The Glossary is made up of three separate pages. Clicking on a letter at the top or the bottom of the page will take you to the first definition starting with that particular letter. Within the Glossary, there are cross references to characters at the same level. Thus when reading the definition of paracytic stomata, you will find direct links to other types of stomata such as anomocytic, cyclocytic, etc. There are also links to definitions at other levels, e.g. to stomata in general, where the basic features of stomata are described, and also to variants of individual stomatal types such as brachyparacytic stomata. We have also built in some synonymy, e.g., Rubiaceous stomata are a synonym of paracytic stomata.
The names of the images of plants may not be up to date - sorry. We did not chose the most spectacular photographs we could find, rather, I tried to select those that showed at least some features on the plant that help characterize the group to which it belongs. All photographs are copyrighted as is evident from the characterizations or the images or the external websites themselves. For photographs of individual species and genera, or simply for more images of families, use Google, but beware of the identifications there!
Website Design: The html code and javascript code for the website were originally generated by hand by Hilary Davis and by using HomeSite © Allaire/Macromedia Corp., and problems dealt with as they appeared. Everything was migrated to Adobe Dreamweaver in March 2015, and a new version of this was installed in November 2019. The general design is a three frame set-up. The top frame contains the "menu," the side frame contains the "quick reference" lists of orders, characters, etc.. The middle frame is the most dynamic and is where most of the pages are displayed. The trees were created by generating a raw dataset in MacClade and drawing the trees in the tree editing window. The tree files were then accessed in PAUP and saved as .pict files to facilitate use in Adobe Illustrator. After polishing the tree images in Illustrator, each tree was saved as a .gif file. Each tree image was made into an image map using a tool in HomeSite, which allows you to select any part of the image (treated as coordinates) and assign a link for those particular coordinates to any place within the website or to any other current website. Note that a number of trees are out of date, so be careful when using them; relationships in the text are more reliable.
ON
THE INTERPRETATION OF THE TREES, TEXT, ABBREVIATIONS, ETC.
[Back to Top]
In the trees in this site I have emphasized mostly nodes with substantial support (e.g. ³80% bootstrap support) that appear after analysis of data from more than one gene; posterior probabilities for a particular node are usually substantially higher than its bootstrap or jacknife values. In a few cases I have been somewhat less cautious, but I have always tried to make it clear where I am treading on thin ice. There are references (not exhaustive) to papers giving support for the relationships suggested here, and these papers may have more resolved trees than those shown, albeit details of these topologies may on occasion have little support. Chase et al (1993, 2000a), Olmstead et al. (1992, 1993, 2000), Olmstead and Graham (2000a, b), Savolainen et al. (2000a, b), D. Soltis et al. (1997, 1998, 2000, 2003, 2005b, 2017), P. Soltis et al. (2000), B. Bremer et al. (2002), Hilu et al. (2003), Jansen et al. (2007), Saarela et al. (2007), Moore et al. (2007, 2010), etc., are invaluable sources for the developing big picture of angiosperm relationships.
If one printed out all the trees in this site and stuck them all together, it might seem as if one had some kind of supertree, however, it is clear from the description of my modus operandi that these trees are hardly a formal supertree. Readers who are interested in the topology of the most parsimonious tree, or relationships suggested by poorly supported nodes, etc., should consult the original literature. PhyloMatic is a another resource to be used.
For an introduction to the interpretation of phylogenetic trees, see Baum and Smith (2012). Krell and Cranston (2004) and Crisp and Cook (2005) and many others have emphasized how careful one must be when interpreting and talking about ladderised trees (see above) in particular and phylogenetic trees in general; the use of the adjective "basal" is especially dangerous (see also D. Soltis et al. 2005b). However, when describing relationships in the context of a branching phylogenetic tree, the use of terms like "high", "low", "basal", etc., to describe aspects of the branching pattern might seem perfectly appropriate, but it is difficult to talk about branching points of a phylogenetic tree; there is no trunk, and no "branch" - one member of a pair of sister clades - can be basal to the other. Although on occasion, I do use the term "basal", this is in the context of a ladderized tree (see above) and refers to the branch in which there has subsequently been less diversification. Thus Equisetum, Amborella, Acorus, Enkianthus, Humbertia, and many other clades can all be called "basal" from this point of view - but note that this has only the topological connotations I have just mentioned, there is certainly no implication that these taxa lack apomorphies or are necessarily "primitive". Indeed, the terms "primitive" and "advanced" are heavily loaded and I have tried not to use them; "plesiomorphic" and its opposite, "derived" and "apomorphic", are the terms to use when talking about individual characters.
In connection with this ladderisation of the trees, comments on particular nodes - whether subtending dichotomies or polytomies - are to be found on the page where the first branch (reading from left to right) at that node is mentioned, and there are cross-links between pages in important cases. Thus comments on the relationships of the magnoliid clade, which involves discussion of the relationships between Chloranthales, monocots, Ceratophyllaceae and eudicots as well, are to be found on the Chloranthales page alone, however, the discussion there is cross-linked to the other pages. However, comments on relationships within the magnoliid group itself are to be found on the Magnoliales page alone.
The information at the beginning of each page is to be read as if you were following along from the base of the embryophyte tree to the branch in which you are interested. Thus each page starts off with a characterisation of the common ancestor of all embryophytes followed by characterisations of all nodes leading up to your branch. Of course, features mentioned more basally in the tree (earlier on that page) may change, perhaps even reverse. Thus at the node [AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]] you will find "ethereal oils +" - this part of the tree seems to be where that feature evolved. However, the apomorphies of [CERATOPHYLLALES + EUDICOTS] and of all monocots minus Acorales include "ethereal oils 0". Ethereal oils have also subsequently been reacquired several times, as in Zingiberaceae and within Lamiaceae.
The families and orders are those recognized by the Angiosperm Phylogeny Group (A.P.G. IV 2016), any differences reflecting subsequent work (see individual pages). Families are grouped within orders as far as possible according to their phylogenetic relationships. Trees showing relationships within many orders and a few of the larger families are included; they may have been cobbled together from more than one study, and I may indicate the level of support for the nodes on the trees. Subfamilies or tribes are numbered sequentially within each family. Although their characterisations may be only imperfectly worked out, they help clarify which characters really are potential family-level apomorphies and which characterise only parts of the family, speciose though those parts may be. The importance of having infra-familial variation pegged to its correct level is evident in, for example, Annonaceae, Convolvulaceae and Fabaceae. Paraphyletic groups are clearly flagged as being such, e.g. "Caesalpinioideae" (paraphyletic) vs Faboideae (monophyletic) - although there is now a particular circumscription of Caesalpinioideae which is monophyletic that can be used (see the Legume Phylogeny Working Group 2017). Further details of relationships, etc., can be found in the papers cited, which should always be consulted.
For the authorities of the names of subfamilies, families, orders, etc., I have relied largely on Reveal's listings (Reveal 2001 onwards), Hoogland and Reveal (2005) and Thorne (with Reveal) (2007). However, the authors of some names as given here may be incorrect since bibliographic work that affects the names of authorities proceeds apace. Synonymy is as complete as I can get it at the familial level and above.
Features are mentioned in the characterisations roughly following the order in the discussion of the characters on the "Characters" page. Possible apomorphies are indicated, but it will be abundantly clear that fixing characters to nodes is no easy task. For some characters (an example is leaf ptyxis) there is little easily-accessible basic information and the terms used to describe it in the literature are not used consistently. It is not always recorded in the characterisations, although its coverage is slowly improving; the same is true for the pattern of petiole vasculature, presence of pericyclic fibers in the stem, etc.; I had initially not considered such characters to be particularly useful (and this indeed may be true of petiole anatomy). Gaps in data are often indicated, but for many features confidently included in a characterisation, e.g. micropyle type, it should not be forgotten that this may be known for very few taxa. The contraction P stands for perianth, T for tepals, K for calyx, C for corolla, A for androecium, and G for gynoecium. "#<" means equal to or more than, "#>" means equal to or fewer than. "Many" means that there are more than fifteen or so parts. Parentheses in characterisations of clades above the level of family denote characters that are scattered throughout that clade, being found in several, but not all, terminal taxa but in no obvious pattern. Examples are septal nectaries and cuticle waxes in monocots, N-fixing in part of the eurosid I clade, and iridoids in asterids. Parentheses in characterizations of terminal taxa refer (as in conventional descriptions) to uncommon features. Square brackets enclose explanations or glosses of the feature described. When a generic name is placed in square brackets after mention of a feature, this emphasized that the feature is known from/I know of it from only that one genus. However, square brackets surrounding carpel number means that the carpels are connate, at least as to their ovule-bearing parts. A line under the number of carpels means that the ovary is superior. A fuller list of abbreviations, etc., used may be found under "Abbreviations" on the top of the left pane.
Following familial and subfamilial characterisations are two figures, the approximate number of accepted genera and species in the group. I mention most genera with 50 or more species and estimate total numbers of species and genera in families; much of this information has been taken from Mabberley's excellent The Plant Book (e.g. Mabberley 2008, 2017) and from accounts in Kubitzki (1993 onwards). A list of included genera and their synonyms can usually be found by clicking on "list". Coverage of the synonymy is variable, and keeping up-to-date on new generic names is impossible. In general, the patient user will be able to answer most questions about the authorities of generic names, commonly-used synonyms, etc.. There are also some links to images.
General distribution is indicated, and there are some hundreds of distribution maps; these are amended as more information becomes available, and there are yet others in process. These give only overall indications of distributions that are unaffected by recent human activities - distributions on oceanic islands in particular are usually very incomplete. In many cases the original outline maps on which many maps here are based have had to been amended, for instance, as substantial holes in the distributions of many apparently widely-ranging families have become apparent - indeed, it has proved surprisingly difficult to find/make accurate maps. Major resources include Hultén and Fries (1986: see also Hultén's earlier work), van Steenis (1963), van Steenis and van Balgooy (1966), van Balgooy (1975, 1984, 1993), Meusel and his collaborators (Meusel et al. 1965, 1978, 1992), and Quian and Ricklefs (2004: see electronic appendix, corrections of the often rather vague maps in Heywood [1978]); invaluable maps for tropical Africa are appearing, see Lebrun and Stork (2003 onwards). Some maps in Heywood et al. (2007) have been used as a last resort. There are various online resouces, including the excellent FloraBase, for the flora of West Australia, and Australia's Virtual Herbarium, for the flora of Australia as a whole; BONAP's Taxonomic Data Center for the north American flora, also provides a useful check on the distributions of plants given in general maps (and also much else). Maps generated by searches on GBIF and Tropicos can be helpful, but beware of records from cultivated plants and aliens. I have included the distributions of fossils when they clarify the distributions of extant taxa. Sources for the maps are indicated.
Following most families and a very few orders is a brief paragraph with features that may help in their recognition; the terms used here may not be perfectly "correct" botanically. Some of these features are taken from LaFrankie (2011), while Utteridge and Bramley (2014) suggest how the commoner tropical families can best be recognized.
There are a number of sections with subheadings following the family accounts (and also at other nodes when relevant), and most of these were introduced starting in late 2008. Age refers to crown-group ages, and these are placed at the appropriate node. In several cases there are substantially different age estimates for the same event, so treat these dates with particular caution. Evolution includes discussion on diversification and biogeography (Diversification & Distribution), comments on mycorrhizae and photosynthetic variants, etc. (Ecology & Physiology), information about pollination and disseminule distribution (Pollination Biology & Seed Dispersal), herbivory and galling (Plant/Animal Interactions), fungal and bacterial associations in general (Plant-Bacterial/Fungal Associations), interesting Vegetative Variation, Genes & Genomes, and plants with particular Economic Importance. Parts of these sections have also found their way to the section on the evolution and diversification of the angiosperms.
In Chemistry, Morphology, etc. interesting variation within the family is summarized and there are references to sources of information that have been used but are not mentioned elsewhere in the family notes; there are additional general references in "Characters". Remember that many of the older "family" and "order" references include information about other groups some of which would not now be considered at all close. Phylogeny includes summaries of major works bearing on our current ideas of phylogentic relationships in the family. Classification includes references to the infrafamilial classification followed here, and there is often some discussion about generic limits. Earlier ideas of relationships (see Previous Relationships) includes largely some mention of some suggestions of Cronquist (1981) and Takhtajan (1997), but only because their work is still used. Finding out who was "first" in suggesting a particular relationship is no goal of these pages, the more so since what is often more interesting is not that a particular suggestion was made, but exactly why it was made. Trivia needs no explanation. The "Bibliography" may still have holes.
In the "Glossary" there are definitions of the terms commonly used in the site and some other terms that may be encountered, representative chemical formulae, etc.. Definitions as far as possible follow current usage, rather than etymology or original definitions (see, e.g. Rickett 1954-56). Building this glossary has forced me to confront head-on such problems as the plethora of terms used to describe inflorescences (in part resolvable, I think, providing one does not attempt too much detail) and fruits (more difficult). Botany needs a common language, and for this the botanical community as a whole needs to agree on definitions. A goal of botanical terminology must be to simplify the terms we use and to try and reduce the number of synonyms in common use.
!!IMPORTANT
- WARNING TO ALL USERS!!
[Back to Top]
All clades are hypotheses of relationships, and as hypotheses they may be overturned. Even though I have for the most part been conservative, changes in our ideas of relationships, and hence in the clades we talk about, are still likely in places. Relationships of a few taxa are still largely unknown or only partly known, and there will turn out to be a few seriously misplaced genera. Thus some changes are to be expected, but change is neither a defect of cladistics nor a necessary consequence of the use of molecular data.
A very important issue for all morphological studies, particularly at this level, is the documentation of variation within a character and the justification of the states that are used to describe this variation. As I often mention, the states used have frequently been inadequately justified, explicit criteria for their delimitation not having been presented. Character states that lack justification may or may not compromise phylogenetic analyses, but there is certainly little reason to talk about their evolution (Stevens 2000 for literature; see also above). To quote Heywood (1973: p. 311) slightly out of context, "Systematics can be likened to a mincing machine into which data of all sorts is fed and processed to form a series of sausages ... The basic recipe for the construction of these "sausages" is usually secret, yet it is such encapsulated pieces of information that we have to work with and communicate with." In general, character states as conventionally delimited may not be as helpful to us as we might like or assume.
For many characters in the characterizations our basic knowledge is often incomplete, and certainly sampling within families is often inadequate. However, I have included less information about chemical, wood anatomical, palynological, and embryological characters than I might. This is because for chemical characters (for example), my basic knowledge needs improving, while for some of the other more cryptic characters the sampling may be poor. But equally importantly, these notes began as teaching notes, and I judged that there was more than enough for the student or even the teacher to be going on with. An ideal system would include much more detail, the user ignoring anything irrelevant for the purpose at hand.
That I mention some features as not being known is in many cases simply an expression of my ignorance.
This site is updated on a more or less continuous basis. I teach plant families at least every other year and that is a particularly good time for finding areas that need attention.
I am always more than grateful if you can let me know of any references in the text that lack citations, errors of omission or commission, etc. - peter.stevens@mobot.org should find me.
Spelling is erratic and somewhat mid-Atlantic; grammar is little better. All errors are mine.
HISTORY OF THE SITE
[Back to Top]
These notes began as I taught Biology 103, the basic plant families course, at Harvard, taking over from Carroll E. Wood, Jr. (an impossible task). They became more wide-ranging when I twice taught OTS "Tropical Plant Systematics", and when I attempted to deal with all families in a one-semester, graduate-level course at Harvard. This last was a failure; we did not get to the monocots. However, the time and energy spent in assembling material for each class from all corners of the herbarium convinced me like nothing else could have that the modified Englerian system the herbarium then followed was didactically a disaster. Finally, I have long been interested in sight identification of herbarium material; unidentified specimens have to be brought into the system somehow. I have done this intermittently, but as frequently as I could, over the years, often with students and others, and this has been invaluable both in reviewing characters and also in learning more.
The first version of the website (version 2) that was made generally known included literature that appeared as late as early August, 2001; the Introduction to this version is dated the 13th of July 2001. Version 2 benefited from teaching again in Costa Rica. This time I was with InBio personnel, and I learned perhaps more than I taught; a course given at the University of Campinas, in Brasil, was similarly stimulating.
What happens with these pages depends on how useful they are found to be, or can be made to be. Their goal is to help in teaching, although I find them also a useful research tool in that they help direct one's attention to interesting characters and to taxa that are poorly understood. They are not intended to compete with other web sites that depict the tree of life, etc.
Filling in gaps in the literature continues to improve family characterisations; three character systems in particular, chemistry, wood anatomy and palynology, clearly need more attention. We will continue to build in more links to photographs and to other web sites, particularly those that focus on families. Adding good diagrams showing the basic floral morphology of each family would be very helpful (see Ronse DeCraene 2010 for recent work on floral diagrams; Eichler 1875-1878 remains invaluable), and simple floral formulae are being added, although complex formulae can be difficult to understand (see Prenner et al. 2010; Simpson 2019; Nuraliev et al. 2019 for other symbols that might be useful).
Last, but certainly not least, I thank Kobinah Abdul-Salim, Stuart Davies, Diane Ferguson, Rick Ree, George Weiblen and Barbara Whitlock for providing the impetus to start this. Students at courses at Harvard University, Massachusetts, InBio, Costa Rica, the University of Campinas, Brasil, and the University of Missouri, St Louis and Missouri Botanical Garden have worked with successive versions of the hierarchy, uncovering flaws as they did do; to them my thanks for their enthusiasm and tolerance. Librarians at Harvard University and the Missouri Botanical Garden have been unfailingly helpful in finding sometimes obscure references. A few sections have been read by colleagues who are thanked individually in the text. Many people have sent in comments and/or noticed mistakes; my thanks to you all, although I have not mentioned you individually unless you have caught particularly egregious errors! Mark Olson, Beth Owen and Bob Magill helped to set up the site. Finally, to say that I am extremely grateful to Hilary Davis for building the site, adding the links to the Gentry photographs of the Missouri Botanical Garden, and generally helping things along, is an understatement.
P. F. Stevens, 13 (Friday) July 2001 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 2.
I inexcusably forgot to thank Dr Barry Hammel (Costa Rica) and Drs Maria Amaral and Volker Bittrich (Campinas, Brasil) for their invitations to teach/learn from InBio personnel and University of Campinas staff and students respectively. For version 3, many have been kind in either giving us their photographs or allowing us to make direct links to their photographs: Atlas of Florida Vascular Plants, Arizona State University Herbarium, Australian National Botanic Gardens, David Boufford - Biodiversity of the Hengduan Mountains China, Mark Brand - UConn Plant Database, CalPhotos, Connecticut Botanical Society, Michael Dillon - Andean Botanical Information System, Michael A. Dirr, Murray Fagg - Australian National Botanic Gardens, Gardenweek.org, Christine Howells - Australian Plants Society Tasmania, Kelly Irvin - International Bulb Society, Don Les, John Maunder - Provincial Museum of Newfoundland and Labrador, Kate McCombs - Christchurch City Council New Zealand, Andrew Morgan - University of Tasmania, Clinton Morse - University of Connecticut Greenhouse, Mount Tomah Botanic Garden Australia, Lytton J. Musselman, Dan Nickrent - Parasitic Plant Connection, Plant of the Week - Smithsonian Institution, James Reveal, George Schatz - Madagascar Conspectus Website, Thomas Schöpke, Tim Stephens - University of California Santa Cruz, Kurt Stüber, Jim Mann Taylor, Texas A&M Bioinformatics Working Group Vascular Plant Gallery, Brian Walters - Australian Society for Growing Native Australian Plants, Len Webb Ecological Images Collection, Western Australia Seagrass Website, Steven J. Wolf, University of Wisconsin Plant Systematics Collection. Gerald Carr and Heinz Schneider have been particularly generous in making available many magnificent photographs; these are but some that can be found at the sites that the two maintain (Vascular Plant Families Image Gallery and Botanical Image Database, respectively). Dave Boufford has caught many errors throughout the site, as has Paul von Rijckevorsel. I am again grateful to students at the University of Missouri at St Louis, Washington University, and St Louis University, and to staff and visitors at the Missouri Botanical Garden. And once again, my sincere thanks to Hilary Davis, to whom the improvements of the site are due.
P. F. Stevens, 15 May 2002 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 3.
Once again, the normal list of suspects is to be thanked. Many people have written in with comments that, whether apparently trivial or cosmic, I have tried to take into account. I remain grateful to students at the University of Missouri at St Louis, Washington University, and St Louis University, and to staff and visitors at the Missouri Botanical Garden. Hilary Davis continues to improve the site.
P. F. Stevens, 2 May 2003 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 4.
Version 4 was archived on May 1. I again thank students at the University of Missouri at St Louis, Washington University, and St Louis University, and all who have caught mistakes, suggested additions, etc. Hilary Davis has worked hard on the site, especially the glossary, and she plans further improvements.
P. F. Stevens, 5 May 2004 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 5.
Version 5 is being archived today, Sunday May 22. All those who have caught mistakes and made suggestions are gratefully thanked, and Hilary Davis is particularly thanked for developing the protocol for making the maps. She has also developed a protocol for integrating illustrations of critical characters with the text that we will try and implement this next year.
P. F. Stevens, 22 May 2005 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 6.
Version 6 was archived on Sunday May 21. All those who have caught mistakes and made suggestions are gratefully thanked, and Andrew Ford has been particularly helpful. The library staff at the Missouri Botanical Garden have patiently dealt with a positive barrage of queries this last year, and I thank Victoria McMichael and Mary Stiffler in particular for their tolerance. Hilary Davies continues to help developing the site.
P. F. Stevens, 22 May 2006 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 7.
Version 7 was archived on Sunday, June 3. I am particularly grateful to students in my plant families course for making this last term so stimulating, and spurring the development of a new section of the site, the "Student" section, which Hilary Davis put up just prior to archiving. Again, the library staff have been swamped with requests, and again corrections and additions have been suggested; I thank all who have been involved in this project, no matter how apparently peripherally.
P. F. Stevens, 8 June 2007 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 8.
Version 8 was archived on Friday, May 30th. The "Student" section has occupied my energies for much of this year. I thank Felipe Zapata for bringing literature to my attention, making suggestions, etc., Hilary Davis for continuing to persevere with this project, and all who have suggested additions and corrections.
P. F. Stevens, Monday, May 26th, 2008 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 9.
Version 9 was archived on Friday, June 19th. In connection with suggestions made by Amy Zanne and Cam Webb, I have been working at making the discussion more rational, introducing a number of subheadings which I hope will make it easier to navigate. Felipe Zapata and Hilary Davis have continued to help with this project, and I thank all of you who have sent me additions and corrections.
P. F. Stevens, Sunday, June 21st, 2009 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 10.
I have just realised that I omitted to say that version 10 was archived on on July 7th; the year has been very hectic. I am not as far along with the reorganization of the site as I wanted to be, but hope springs eternal. Cam Webb, Amy Zanne, Felipe Zapata and Hilary Davis continue with the project, and again I thank all of you who have sent me additions and corrections; the "small" errors often point out larger problems.
P. F. Stevens, Saturday, November 20th(!), 2010 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 11.
Version 11 was archived on May 21st. Unfortunately, this year was no better than the last... Thanks Cam, Amy, Felipe and Hilary for continuing to directly or indirectly prod, and to all for corrections; trees will have to be updated in the very near future!
P. F. Stevens, Saturday, May 28, 2011 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 12.
Version 12 was archived on July 25th. The whole site is in the process of being reworked, and this has exposed the consequences of adding bits of information sporadically over a long period of time... In addition, with the help of Cam Webb the apomorphies - many of which have been added this past year - are being given xml tags, and these will shortly be extended to the discussion, the nodes, etc. Thanks to Amy, Felipe and Hilary for continuing help, the staff of the library at the Garden who have deat with even more of a barrage of requests than usual, and to Volker Bittrich and to all the rest of you for comments and corrections.
P. F. Stevens, Tuesday, August 1, 2012 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 13.
Version 13 was archived on September 26th. The reworking of the site has continued - apomorphies have now been flagged throughout the site, and dates for clades, which were in a considerable mess, are being cleaned up. Given the reorganisation of the site over the last couple of years, the "Student" section has been removed, as has the "Apomorphies" section (it is now redundant) and the "Statistics" section (that was simply one more area to keep up to date, and anyhow, I had largely ignored it).
P. F. Stevens, Tuesday, September 28, 2013 - University of Missouri, St Louis, and Missouri Botanical Garden.
Version 14.
Getting there.... Dates for clades are now largely cleaned up, but most must be seriously inaccurate. The next push must be to clean up the trees. Readers have made many useful comments, and Felipe Zapata in particular has helped with literature.
P. F. Stevens, July 4, 2017 - University of Missouri, St Louis, and Missouri Botanical Garden.