EMBRYOPSIDA Pirani & Prado
Gametophyte dominant, independent, multicellular, initially ±globular, not motile, branched; showing gravitropism; glycolate oxidase +, glycolate metabolism in leaf peroxisomes [glyoxysomes], acquisition of phenylalanine lysase* [PAL], flavonoid synthesis*, microbial terpene synthase-like genes +, triterpenoids produced by CYP716 enzymes, CYP73 and phenylpropanoid metabolism [development of phenolic network], xyloglucans in primary cell wall, side chains charged; plant poikilohydrous [protoplasm dessication tolerant], ectohydrous [free water outside plant physiologically important]; thalloid, leafy, with single-celled apical meristem, tissues little differentiated, rhizoids +, unicellular; chloroplasts several per cell, pyrenoids 0; centrioles/centrosomes in vegetative cells 0, microtubules with γ-tubulin along their lengths [?here], interphase microtubules form hoop-like system; metaphase spindle anastral, predictive preprophase band + [with microtubules and F-actin; where new cell wall will form], phragmoplast + [cell wall deposition centrifugal, from around the anaphase spindle], plasmodesmata +; antheridia and archegonia +, jacketed*, surficial; blepharoplast +, centrioles develop de novo, bicentriole pair coaxial, separate at midpoint, centrioles rotate, associated with basal bodies of cilia, multilayered structure + [4 layers: L1, L4, tubules; L2, L3, short vertical lamellae] (0), spline + [tubules from L1 encircling spermatid], basal body 200-250 nm long, associated with amorphous electron-dense material, microtubules in basal end lacking symmetry, stellate array of filaments in transition zone extended, axonemal cap 0 [microtubules disorganized at apex of cilium]; male gametes [spermatozoids] with a left-handed coil, cilia 2, lateral, asymmetrical; oogamy; sporophyte +*, multicellular, growth 3-dimensional*, cuticle +*, plane of first cell division transverse [with respect to long axis of archegonium/embryo sac], sporangium and upper part of seta developing from epibasal cell [towards the archegonial neck, exoscopic], with at least transient apical cell [?level], initially surrounded by and dependent on gametophyte, placental transfer cells +, in both sporophyte and gametophyte, wall ingrowths develop early; suspensor/foot +, cells at foot tip somewhat haustorial; sporangium +, single, terminal, dehiscence longitudinal; meiosis sporic, monoplastidic, MTOC [= MicroTubule Organizing Centre] associated with plastid, sporocytes 4-lobed, cytokinesis simultaneous, preceding nuclear division, quadripolar microtubule system +; wall development both centripetal and centrifugal, 1000 spores/sporangium, sporopollenin in the spore wall* laid down in association with trilamellar layers [white-line centred lamellae; tripartite lamellae]; plastid transmission maternal; nuclear genome [1C] <1.4 pg, main telomere sequence motif TTTAGGG, KNOX1 and KNOX2 [duplication] and LEAFY genes present, ethylene involved in cell elongation; chloroplast genome with close association between trnLUAA and trnFGAA genes [precursors for starch synthesis], tufA, minD, minE genes moved to nucleus; mitochondrial trnS(gcu) and trnN(guu) genes +.
Many of the bolded characters in the characterization above are apomorphies of more or less inclusive clades of streptophytes along the lineage leading to the embryophytes, not apomorphies of crown-group embryophytes per se.
All groups below are crown groups, nearly all are extant. Characters mentioned are those of the immediate common ancestor of the group, [] contains explanatory material, () features common in clade, exact status unclear.
POLYSPORANGIOPHYTA†
Sporophyte well developed, branched, branching dichotomous, potentially indeterminate; hydroids +; stomata on stem; sporangia several, terminal; spore walls not multilamellate [?here].
II. TRACHEOPHYTA / VASCULAR PLANTS
Sporophyte long lived, cells polyplastidic, photosynthetic red light response, stomata open in response to blue light; plant homoiohydrous [water content of protoplasm relatively stable]; control of leaf hydration passive; plant endohydrous [physiologically important free water inside plant]; PIN[auxin efflux facilitators]-mediated polar auxin transport; (condensed or nonhydrolyzable tannins/proanthocyanidins +); borate cross-linked rhamnogalactan II, xyloglucans with side chains uncharged [?level], in secondary walls of vascular and mechanical tissue; lignins +; roots +, often ≤1 mm across, root hairs and root cap +; stem apex multicellular [several apical initials, no tunica], with cytohistochemical zonation, plasmodesmata formation based on cell lineage; vascular development acropetal, tracheids +, in both protoxylem and metaxylem, G- and S-types; sieve cells + [nucleus degenerating]; endodermis +; stomata numerous, involved in gas exchange; leaves +, vascularized, spirally arranged, blades with mean venation density ca 1.8 mm/mm2 [to 5 mm/mm2], all epidermal cells with chloroplasts; sporangia in strobili, sporangia adaxial, columella 0; tapetum glandular; sporophyte-gametophyte junction lacking dead gametophytic cells, mucilage, ?position of transfer cells; MTOCs not associated with plastids, basal body 350-550 nm long, stellate array in transition region initially joining microtubule triplets; archegonia embedded/sunken [only neck protruding]; embryo suspensor +, shoot apex developing away from micropyle/archegonial neck [from hypobasal cell, endoscopic], root lateral with respect to the longitudinal axis of the embryo [plant homorhizic].
[MONILOPHYTA + LIGNOPHYTA]Sporophyte growth ± monopodial, branching spiral; roots endomycorrhizal [with Glomeromycota], lateral roots +, endogenous; G-type tracheids +, with scalariform-bordered pits; leaves with apical/marginal growth, venation development basipetal, growth determinate; sporangium dehiscence by a single longitudinal slit; cells polyplastidic, MTOCs diffuse, perinuclear, migratory; blepharoplasts +, paired, with electron-dense material, centrioles on periphery, male gametes multiciliate; nuclear genome [1C] 7.6-10 pg [mode]; chloroplast long single copy ca 30kb inversion [from psbM to ycf2]; mitochondrion with loss of 4 genes, absence of numerous group II introns; LITTLE ZIPPER proteins.
LIGNOPHYTA†
Sporophyte woody; stem branching axillary, buds exogenous; lateral root origin from the pericycle; cork cambium + [producing cork abaxially], vascular cambium bifacial [producing phloem abaxially and xylem adaxially].
SEED PLANTS† / SPERMATOPHYTA†
Growth of plant bipolar [plumule/stem and radicle/root independent, roots positively geotropic]; plants heterosporous; megasporangium surrounded by cupule [i.e. = unitegmic ovule, cupule = integument]; pollen lands on ovule; megaspore germination endosporic, female gametophyte initially retained on the plant, free-nuclear/syncytial to start with, walls then coming to surround the individual nuclei, process proceeding centripetally.
EXTANT SEED PLANTS
Plant evergreen; nicotinic acid metabolised to trigonelline, (cyanogenesis via tyrosine pathway); microbial terpene synthase-like genes 0; primary cell walls rich in xyloglucans and/or glucomannans, 25-30% pectin [Type I walls]; lignin chains started by monolignol dimerization [resinols common], particularly with guaiacyl and p-hydroxyphenyl [G + H] units [sinapyl units uncommon, no Maüle reaction]; roots often ≥1 mm across, stele diarch to pentarch, xylem and phloem originating on alternating radii, cork cambium deep seated, gravitropism response fast; stem apical meristem complex [with quiescent centre, etc.], plasmodesma density in SAM 1.6-6.2[mean]/μm2 [interface-specific plasmodesmatal network]; eustele +, protoxylem endarch, endodermis 0; wood homoxylous, tracheids and rays alone, tracheid/tracheid pits circular, bordered; mature sieve tube/cell lacking functioning nucleus, sieve tube plastids with starch grains; phloem fibres +; cork cambium superficial; leaf nodes 1:1, a single trace leaving the vascular sympodium; leaf vascular bundles amphicribral; guard cells the only epidermal cells with chloroplasts, stomatal pore with active opening in response to leaf hydration, control by abscisic acid, metabolic regulation of water use efficiency, etc.; branching by axillary buds, exogenous; prophylls two, lateral; leaves with petiole and lamina, development basipetal, lamina simple; sporangia borne on sporophylls; spores not dormant; microsporophylls aggregated in indeterminate cones/strobili; grains monosulcate, aperture in ana- position [distal], primexine + [involved in exine pattern formation with deposition of sporopollenin from tapetum there], exine and intine homogeneous, exine alveolar/honeycomb; ovules with parietal tissue [= crassinucellate], megaspore tetrad linear, functional megaspore single, chalazal, sporopollenin 0; gametophyte ± wholly dependent on sporophyte, development initially endosporic [apical cell 0, rhizoids 0, etc.]; male gametophyte with tube developing from distal end of grain, male gametes two, developing after pollination, with cell walls; embryo cellular ab initio, suspensor short-minute, embryonic axis straight [shoot and root at opposite ends], primary root/radicle produces taproot [= allorhizic], cotyledons 2; embryo ± dormant; chloroplast ycf2 gene in inverted repeat, trans splicing of five mitochondrial group II introns, rpl6 gene absent; ??whole nuclear genome duplication [ζ/zeta duplication event], 2C genome size (0.71-)1.99(-5.49) pg, two copies of LEAFY gene, PHY gene duplications [three - [BP [A/N + C/O]] - copies], 5.8S and 5S rDNA in separate clusters.
IID. ANGIOSPERMAE / MAGNOLIOPHYTA
Lignans, O-methyl flavonols, dihydroflavonols, triterpenoid oleanane, apigenin and/or luteolin scattered, [cyanogenesis in ANA grade?], lignin also with syringyl units common [G + S lignin, positive Maüle reaction - syringyl:guaiacyl ratio more than 2-2.5:1], hemicelluloses as xyloglucans; root cap meristem closed (open); pith relatively inconspicuous, lateral roots initiated immediately to the side of [when diarch] or opposite xylem poles; epidermis probably originating from inner layer of root cap, trichoblasts [differentiated root hair-forming cells] 0, hypodermis suberised and with Casparian strip [= exodermis]; shoot apex with tunica-corpus construction, tunica 2-layered; starch grains simple; primary cell wall mostly with pectic polysaccharides, poor in mannans; tracheid:tracheid [end wall] plates with scalariform pitting, multiseriate rays +, wood parenchyma +; sieve tubes enucleate, sieve plates with pores (0.1-)0.5-10< µm across, cytoplasm with P-proteins, not occluding pores of plate, companion cell and sieve tube from same mother cell; ?phloem loading/sugar transport; nodes 1:?; dark reversal Pfr → Pr; protoplasm dessication tolerant [plant poikilohydric]; stomata randomly oriented, brachyparacytic [ends of subsidiary cells ± level with ends of guard cells], outer stomatal ledges producing vestibule, reduction in stomatal conductance with increasing CO2 concentration; lamina formed from the primordial leaf apex, margins toothed, development of venation acropetal, overall growth ± diffuse, secondary veins pinnate, fine venation hierarchical-reticulate, (1.7-)4.1(-5.7) mm/mm2, vein endings free; flowers perfect, pedicellate, ± haplomorphic, protogynous; parts free, numbers variable, development centripetal; P = T, petal-like, each with a single trace, outer members not sharply differentiated from the others, not enclosing the floral bud; A many, filament not sharply distinguished from anther, stout, broad, with a single trace, anther introrse, tetrasporangiate, sporangia in two groups of two [dithecal], each theca dehiscing longitudinally by a common slit, ± embedded in the filament, walls with at least outer secondary parietal cells dividing, endothecium +, cells elongated at right angles to long axis of anther; tapetal cells binucleate; microspore mother cells in a block, microsporogenesis successive, walls developing by centripetal furrowing; pollen subspherical, tectum continuous or microperforate, ektexine columellate, endexine restricted to the apertural regions, thin, compact, intine in apertural areas thick, orbicules +, pollenkitt +; nectary 0; carpels present, superior, free, several, spiral, ascidiate [postgenital occlusion by secretion], stylulus at most short [shorter than ovary], hollow, cavity not lined by distinct epidermal layer, stigma ± decurrent, carinal, dry; suprastylar extragynoecial compitum +; ovules few [?1]/carpel, marginal, anatropous, bitegmic, micropyle endostomal, outer integument 2-3 cells across, often largely subdermal in origin, inner integument 2-3 cells across, often dermal in origin, parietal tissue 1-3 cells across, nucellar cap?; megasporocyte single, hypodermal, functional megaspore lacking cuticle; female gametophyte lacking chlorophyll, four-celled [one module, egg and polar nuclei sisters]; ovule not increasing in size between pollination and fertilization; pollen grains bicellular at dispersal, germinating in less than 3 hours, siphonogamy, pollen tube unbranched, growing towards the ovule, between cells, growth rate (ca 10-)80-20,000 µm h-1, tube apex of pectins, wall with callose, lumen with callose plugs, penetration of ovules via micropyle [porogamous], whole process takes ca 18 hours, distance to first ovule 1.1-2.1 mm; male gametophytes tricellular, gametes 2, lacking cell walls, ciliae 0, double fertilization +, ovules aborting unless fertilized; fruit indehiscent, P deciduous; mature seed much larger than fertilized ovule, small [<5 mm long], dry [no sarcotesta], exotestal; endosperm +, ?diploid [one polar nucleus + male gamete], cellular, development heteropolar [first division oblique, micropylar end initially with a single large cell, divisions uniseriate, chalazal cell smaller, divisions in several planes], copious, oily and/or proteinaceous, embryo short [<¼ length of seed]; plastid and mitochondrial transmission maternal; Arabidopsis-type telomeres [(TTTAGGG)n]; nuclear genome [2C] (0.57-)1.45(-3.71) [1 pg = 109 base pairs], ??whole nuclear genome duplication [ε/epsilon event]; ndhB gene 21 codons enlarged at the 5' end, single copy of LEAFY and RPB2 gene, knox genes extensively duplicated [A1-A4], AP1/FUL gene, palaeo AP3 and PI genes [paralogous B-class genes] +, with "DEAER" motif, SEP3/LOFSEP and three copies of the PHY gene, [PHYB [PHYA + PHYC]]; chloroplast IR expansions, chlB, -L, -N, trnP-GGG genes 0.
[NYMPHAEALES [AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]]]: wood fibres +; axial parenchyma diffuse or diffuse-in-aggregates; pollen monosulcate [anasulcate], tectum reticulate-perforate [here?]; ?genome duplication; "DEAER" motif in AP3 and PI genes lost, gaps in these genes.
[AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]]: phloem loading passive, via symplast, plasmodesmata numerous; vessel elements with scalariform perforation plates in primary xylem; essential oils in specialized cells [lamina and P ± pellucid-punctate]; tension wood + [reaction wood: with gelatinous fibres, G-fibres, on adaxial side of branch/stem junction]; anther wall with outer secondary parietal cell layer dividing; tectum reticulate; nucellar cap + [character lost where in eudicots?]; 12BP [4 amino acids] deletion in P1 gene.
[MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]] / MESANGIOSPERMAE: benzylisoquinoline alkaloids +; sesquiterpene synthase subfamily a [TPS-a] [?level], polyacetate derived anthraquinones + [?level]; outer epidermal walls of root elongation zone with cellulose fibrils oriented transverse to root axis; P more or less whorled, 3-merous [?here]; pollen tube growth intra-gynoecial; extragynoecial compitum 0; carpels plicate [?here]; embryo sac monosporic [spore chalazal], 8-celled, bipolar [Polygonum type], antipodal cells persisting; endosperm triploid.
[CERATOPHYLLALES + EUDICOTS]: ethereal oils 0 [or next node up]; fruit dry [very labile].
EUDICOTS: (Myricetin +), asarone 0 [unknown in some groups, + in some asterids]; root epidermis derived from root cap [?Buxaceae, etc.]; (vessel elements with simple perforation plates in primary xylem); nodes 3:3; stomata anomocytic; flowers (dimerous), cyclic; protandry common; K/outer P members with three traces, ("C" +, with a single trace); A ?, filaments fairly slender, anthers basifixed; microsporogenesis simultaneous, pollen tricolpate, apertures in pairs at six points of the young tetrad [Fischer's rule], cleavage centripetal, wall with endexine; G with complete postgenital fusion, stylulus/style solid [?here], short [<2 x length of ovary]; seed coat?; palaeotetraploidy event.
[PROTEALES [TROCHODENDRALES [BUXALES + CORE EUDICOTS]]]: (axial/receptacular nectary +).
[TROCHODENDRALES [BUXALES + CORE EUDICOTS]]: benzylisoquinoline alkaloids 0; euAP3 + TM6 genes [duplication of paleoAP3 gene: B class], mitochondrial rps2 gene lost.
[BUXALES + CORE EUDICOTS]: mitochondrial rps11 gene lost.
CORE EUDICOTS / GUNNERIDAE Back to Main Tree
(Ellagic and gallic acids +); leaf margins serrate; compitum + [one position]; micropyle?; γ/gamma genome duplication [allopolyploidy, 4x x 2x], x = 3 x 7 = 21, 2C genome size (0.79-)1.05(-1.41) pg, PI-dB motif +; small deletion in the 18S ribosomal DNA common.
Note: In all node characterizations, boldface denotes a possible apomorphy, (....) denotes a feature the exact status of which in the clade is uncertain, [....] includes explanatory material; other text lists features found pretty much throughout the clade. However, the precise node to which many characters, particularly the more cryptic ones, should be assigned is unclear. This is partly because homoplasy is very common, in addition, basic information for all too many characters is very incomplete, frequently coming from taxa well embedded in the clade of interest and so making the position of any putative apomorphy uncertain. Then there are the not-so-trivial issues of how character states are delimited and ancestral states are reconstructed (see above).
Age. The crown group age of core eudicots is around 115.5 Ma (Anderson et al. 2005). Other crown group ages are 132-119(-89) Ma in Soltis et al. (2008: a variety of estimates), 114.4 or 115.1 Ma in Magallón and Castillo (2009), (113-)110(-105) Ma (M. J. Moore et al. 2010: 95% HPD), (139-)127, 119(-109) or (127-)121, 117(-107) Ma (Bell et al. 2010), (116-)112(-107) Ma (N. Zhang et al. 2012) and (118.3-)112.9-111.7(-105.3) Ma (Magallón et al. 2013: with temporal constraints); Schneider et al. (2004) offer a range of dates up to over 180 Ma, and another high estimate is ca 171Ma by Z. Wu et al. (2014). Wikström et al. (2003) suggested a crown group age of (131-)127, 116(-111) Ma, Magallón et al. (2015) an age of around 125.1 Ma, ca 118.1 Ma is the age in Naumann et al. (2013), about 117.5 Ma in Tank et al. (2015: Table S1), but only (108-)98(-87) Ma in Murat et al. (2017) and around 130.2-124.4 Ma in Zeng et al. (2017).
The age of stem core eudicots has been estimated as some 120-116 Ma (Anderson et al. 2005). Other stem group ages are 121.2 or 121.9 Ma (relaxed and constrained penalized likelihood: Magallón & Castillo 2009), while Wikström et al. (2003) suggested a stem age about (140-)135, 123(-118) Ma - c.f. sister groups. See also below for the age of the palaeohexaploidy event that probably occured somewhere along the core eudicot stem.
Evolution: Divergence & Distribution. Vekemans et al. (2012) emphasized that the γ genome duplication event (see below) occurred ca 7 Ma before the divergence of Gunnerales and other core eudicots; there was a lag between duplication and subsequent divergence. Indeed, if one tries to understand any link between this event and diversification, the lag may be somewhat greater, since although Gunnerales may be chemically like other core eudicots, they are not particularly similar florally, and in any event they hardly represent a particularly speciose clade (Mayrose et al. 2011; Schranz et al. 2012 for lags between genome duplication/polyploidy and diversification). There seems not to have been immediate subsequent global diversification, at least in the clade that includes Coffea and Cephalotus, i.e. in the bulk of the core eudicots (Fukushima et al. 2017). However, Tank et al. (2015) put the increase in diversification rate possibly associated with the duplication at the Pentapetalae node, indeed, L. Zhang et al. (2020) suggest that the duplication may have been in some way involved in e functional diversification of floral organ developmental regulators, and hence in the evolution of the flower - discussed further elsewhere. Puttick et al. (2015) thought that of the seven genome duplications that they examined (including Brassicaceae, Asteraceae, angiosperms, seed plants), only here were there accelerated rates both for genome size evolution and speciation. However, understanding patterns of diversification is made more difficult by the suggestion that the γ genome duplication event is in fact two events, the first of which is at the base of the eudicots (Aköz & Nordborg 2019) and perhaps around 10-20 Ma before the second (see below, furthermore, relationships at the base of the Pentapetalae are unclear - an understatement. Interestingly, Y. Wang et al. (2016) found that a large number of genes involved in the γ genome event quite quickly became dispersed throughout the genome, perhaps by chromosome breakage and reassembly - note, these are a different class of genes from those that make up normal steady-state dispersed duplication events. Moreover, these relocated γ duplicates showed relaxed purifying selection and are significantly enriched in genes with essential functions compared to other γ duplicate genes; such a widespread relocation of genes following a duplication event is unusual (Wang et al. 2016). For duplication(s) around here, see also Zwaenepoel and Van de Peer (2019).
The floral morphology of Gunnerales is more like that of Buxales, etc., than that of other core eudicots. That is, it is more like taxa on more basal eudicot branches and so it is apparently plesiomorphic (e.g. D. Soltis et al. 2003a; Doust & Stevens 2005; Kubitzki 2006a). Wanntorp and Ronse De Craene (2005) and Ronse De Craene and Wanntorp (2006) also note that the morphology of Gunnerales flowers cannot be directly related to that of the pentamerous core eudicots, the floral morphology of the former being shaped by the exigencies of wind pollination (see also Ronse de Craene & Brockington 2013). Wanntorp and Ronse De Craene (2005) observed that three successive floral whorls may be opposite each other, and this is a feature of some Ranunculales-Berberidaceae, Proteales-Sabiaceae, etc., but not of core eudicots. Indeed, the flowers cannot readily be related to those of other core eudicots given our current knowledge of floral morphology and development, even if the duplication of a number of genes important in determining the identity of floral parts (AP3, AP1, SEP, AG) may be connected with the γ duplication event (Jiao et al. 2012; see below). For discussion of the "typical" pentapetalous core eudicot flower, see the Dilleniales page.
Chemically Gunnerales are similar to Pentapetalae in that they have ellagic acid (e.g. Soltis et al. 2005b), and general molecular data link them closely with that clade. Some perhaps important gene duplications may have occurred around here or somewhat earlier (see the Trochodendrales page), and it is not known if Gunnerales have the euAP3 gene, etc. (e.g. Hernández-Hernández et al. 2006; Gloppato & Dornelas 2018).
Seago et al. (2021) floated a suggestion that the distinctive anatomy and more or less aquatic habitats of Gunnera might provide clues as to the morphology and habitats of basal core eudicots; interestingly, G. herteri, sister to the rest of the family, has a rather less "extreme" anatomy compared with some other members of the genus. However, there was no mention of Myrothamnaceae, which, using srimilar arguments, should provide similar clues... Rather, it would seem that both Myrothamnaceae and Gunneraceae are likely to be more or less highly derived.
For possible ecophysiological and morphological changes here associated with the γ duplication, and in more or less contemporaneous duplications in Magnoliales + Laurales, Nymphaeales, and in the monocots, perhaps at the [Asparagales + commelinds] node, see elsewhere.
For general information on core eudicot diversification, see Magallón et al. (1999); most of the estimates of percentage diversity of clades are taken from this work. The diversification rates of many of the clades are higher than those in other angiosperms (Magallón & Sanderson 2001).
Pollination Biology & Seed Dispersal. Compitum presence can perhaps be pegged as to this node (?a key innovation?) given that it is found in Gunneraceae, the rosids, and the extended asterid clade, although it is absent in Myrothamnaceae and Dilleniaceae (c.f. Endress 2011).
Genes & Genomes. It is suggested that x = 7 was the base number in the ancestor of core eudicots, i.e. after the divergence of Buxales and Trochodendrales, x = 21 after the genome triplication, i.e., this is the ancestral core eudicot karyotype; there had been substantial genome rearrangements by the time Vitales diverged from other rosids (Murat et al. 2015b and references, 2017; J. Wang et al. 2017). Indeed, Aköz and Nordborg (2019) thought that rearrangements in Vitales themselves had not been that great. For an estimate of genome size at this node, see Puttick et al. (2015).
The importance of what was thought to be a palaeohexaploidy event (the γ triplication, or core-eudicot common hexaploidization - ECH), was first suggested in work on Vitales (Jaillon, Aury et al. 2007). At first sight it appeared that there had been a gene duplication, possibly because of hybridization, within the Vitis lineage itself, and that this brought the Vitis genome more into line with that of other rosids (Velasco et al. 2007). Nevertheless, Freeling et al. (2008: Carica papaya genome included) thought that most rosids, i.e. the node [Vitales + rosids s. str], were palaeohexaploids, the Atg (γ) event. Evidence from genome colinearity suggested that this palaeohexaploidy event had also occured in the ancestor of the asterid Coffea (Cenci et al. 2010; see also the Potato Sequencing Consortium 2011: c.f. monocots and eudicots), and Jiao et al. (2012) placed this event before the split of the asterids and rosids but after the divergence of Ranunculales - see also Tang et al. (2008a, b), Diaz-Riquelme et (al. 2009), Barker et al. (2009), Abrouk et al. (2010), Severin et al. (2011), Dohm et al. (2011), Zheng et al. (2013), Zumajo-Cardona et al. (2017) and Gao et al. (2018) for this genome triplication, whether or not also involving asterids. In any event, evidence of large genome duplications may be lost, as in Fragaria vesca (Rosaceae) (Shulaev et al. 2010).
It has been suggested that this palaeohexaploidy event occurred in the immediate ancestor of this node, the core eudicot node (Jiao et al. 2012; esp. Vekemans et al. 2012). Severin et al. (2011) offered a spread of 240-130 Ma for the age of this event, Vekemans et al. (2012) narrowed the age estimate to (121.9-)120.4(-118.9) or (122.7-)120.05(-117.4) Ma, depending on the method used, while Jiao et al. (2012) dated the duplications to around 117 Mya; 138-11 Ma is the estimate in Murat et al. (2015b), ca 130 Ma that in J. Wang et al. (2017), ca 110 Ma that in L. Zhang et al. (2020) and 129.4-103.7 Ma in X. Guo et al. (2021).
Note that Freeling et al. (2008) had found that two of the three genomes involved had lost significantly more ancestral genes than had the other, perhaps a little odd. Similarly, Chanderbali et al. (2016b; see also Ming et al. 2013) suggested that only around half the gene duplications associated with this event were to be placed here, the other half being earlier, perhaps being placed at the [Trochodendrales, Buxales, core eudicot] node. Did the palaeohexaploidy involve an ancestor of Nelumbo and another species that had branched off at about the same time (Ming et al. 2013)? Other earlier work suggested that there were two duplications, for instance, Malacarne et al. (2012) proposed that the component genomes had 40-60 Ma to evolve independently (see also Jiao & Paterson 2013). Indeed, recent work suggests that there was an allopolyploidy event at the core eudicot node involving 4x and 2x plants that was preceded by an earlier tetraploidy event at the eudicot node (Aköz & Nordborg 2019). However, P.-L. liu et al (2020) talk about duplications in the Aquilegia, Lotus, Vitis and Trochodendraceae (two) lineages, but these seem not to be related to the major duplications that are being discussed. Shi and Chen (2020) also suggest that the supposed eudicot duplication was in fact perhaps within Ranunculaceae-Aquilegia and closer than Papaver (see also Y. Liu et al. 2021) - but see reply by Aköz and Nordborg (2020). P.-L. Liu et al (2020) also dismissed any global importance of the duplications they discussed in Aquilegia, Lotus and Trochodendrales, eudicots were not involved in this major duplication, rather, it happened at around the core eudicot node.
Given earlier genome duplications (all angiosperms, all seed plants), by the time of the evolution of genera like Brassica, Arabidopsis and Gossypium in which there have been several more local duplications, the genome must have duplicated many, many times... (e.g. Paterson et al. 2012). Furthermore, how genomes have subsequently been rearranged varies greatly, Aköz and Nordborg (2019) suggesting that there were only 3 post-γ rearrangements in Vitis, while Badouin et al. (2017) estimated 50 times as many, some 150, in Helianthus.
Whether or not the chloroplast rps2 and rps11 genes occur in Gunnerales is apparently unknown, but the first is absent in Buxales and Trochodendrales and the second in Buxales alone (Adams et al. 2002b); they are probably lost slightly lower down on the tree (as are benzylisoquinoline alkaloids).
Chemistry, Morphology, etc.. Wanntorp and Ronse De Craene (2005) and Ronse De Craene and Wanntorp (2006) note that the morphology of Gunnerales flowers cannot be directly related to that of the pentamerous core eudicots, the floral morphology of the former being shaped by the exigencies of wind pollination. Wanntorp and Ronse De Craene (2005) also note that three successive floral whorls may be opposite each other, and this is a feature e.g. of some Ranunculales like Berberidaceae, Sabiaceae, etc., but not of core eudicots. Indeed, the flowers cannot be considered as being readily derived from those of other core eudicots given our current knowledge of floral morphology and development, rather, they seem more similar to some of those scattered in other eudicot clades basal to Gunnerales. For discussion of the "typical" core eudicot flower, see the Pentapetalae page.
Phylogeny. This clade is strongly supported, e.g. Chase et al. (1993), D. Soltis et al. (1997, 1999, 2003a), Hoot et al. (1998), Nandi et al. (1998), D. Soltis et al. (2003a: four genes), S. Kim et al. (2004), Zhu et al. (2007), Qiu et al. (2010: support weak), and Z. Wu et al. (2014), although indistinguishable from other core eudicots in (e.g.) P. Soltis et al. (1999). However, in some earlier studies the exact position of Gunnerales was unclear (e.g. Chase et al. 1993; Morgan & Soltis 1993), but quite strong support for a position sister to the rest of the core eudicots was provided by Senters et al. (2000: Gunneraceae alone sampled), rather weaker support by Hilu et al. (2001). Zhu et al. (2007) found a weakly supported [Gunnera + Dillenia] clade sister to other core eudicots; see also Pentapetalae for discussion of relationships immediately above Gunnerales on the tree.
Classification. Gunnerales were excluded from the core eudicots in earlier versions of this site (pre Version 7; c.f. A.P.G. I and II 1999, 2003) because of their apparently largely plesiomorphic floral morphology. However, the limits of core eudicots have been adjusted to conform with those now generally used; classifications based on the one topology should be the same.
GUNNERALES Reveal - Main Tree.
Ellagic acid +; vessel elements?; sieve tube plastids with protein crystalloids and starch; pith with sclerenchymatous diagrams; lamina margins with hydathodal teeth, secondary veins palmate; plants dioecious; inflorescences with terminal flowers; flowers small [<5 mm across]; A latrorse; stigma at most weakly secretory; seed coat? - 2 families, 2 genera, 42-52 species.
Includes Gunneraceae, Myrothamnaceae.
Note: In all node characterizations, boldface denotes a possible apomorphy, (....) denotes a feature the exact status of which in the clade is uncertain, [....] includes explanatory material; other text lists features found pretty much throughout the clade. However, the precise node to which many characters, particularly the more cryptic ones, should be assigned is unclear. This is partly because homoplasy is very common, in addition, basic information for all too many characters is very incomplete, frequently coming from taxa well embedded in the clade of interest and so making the position of any putative apomorphy uncertain. Then there are the not-so-trivial issues of how character states are delimited and ancestral states are reconstructed (see above).
Age. Estimates of the age of crown group Gunnerales are (123-)118, 108(-103) Ma (Wikström et al. 2001), about 104.6 Ma (Magallón et al. 2015), 90-55 Ma (Anderson et al. 2005), (132-)102, 95(-64) Ma (Bell et al. 2010), ca 99.8 Ma (Tank et al. 2015: Table S2), or ca 77 Ma (Magallón et al. 2013).
The fossil Spanomera mauldinensis described from east North American deposits laid down 113-98 Ma in the Albian (Drinnan et al. 1991) is perhaps close to Buxaceae or Myrothamnaceae (see also Crepet et al. 2004; Doyle & Endress 2010; Doyle & Upchurch 2014). Recent work suggests that it is in the Buxales-Gunnerales area, whether in their stem or crown groups or in an independant lineage (Schönenberger et al. 2020).
Evolution: Divergence & Distribution. González and Bello (2009) suggest possible apomorphies for the order, including the presence of stipules, but see below for a possible interpretation of the stipules of Gunnera.
Chemistry, Morphology, etc.. In Gunneraceae, hydathodes are well developed and mucilage or possibly resinous lacquer is secreted, in Myrothamnaceae, hydathodes are poorly developed (but see Drennan et al. 2009) and the plant secretes resin.
Both families have flowers without much of a perianth, although the plesiomorphic condition for the order may be to have some kind of perianth. Details of pollen morphology (e.g. c.f. Zavada & Dilcher 1986; Wanntorp et al. 2004), etc., differ.
Classification. There is an option of including Myrothamnaceae in Gunneraceae in A.P.G. II (2003), both being small and monogeneric families, but they are so different in appearance that it seems best to keep them separate (see Wilkinson 2000 for a table of differences).
Previous Relationships. This is a rather surprising group. Gunneraceae and Myrothamnaceae look very different; one is an often gigantic mesophytic herb, the other a resurrection shrub of arid habitats. Gunneraceae have often been associated with Haloragaceae (e.g. Cronquist 1981), also with an inferior ovary and reduced flowers, but in the latter the stamens are as many as the sepals, and opposite them, the gynoecium is multilocular, with one ovule/loculus, etc. - see Saxifragales. Gunneraceae were included in Saxifraganae-Rosidae by Takhtajan (1997). Fuller and Hickey (2005), examining details of leaf architecture, etc., suggested that Gunneraceae were best associated with the herbaceous Saxifragaceae, which, they thought, "provided a more coherent understanding of the evolutionarty history of Gunnera" (ibid. p. 295) than did suggestions of a link with Myrothamnaceae, but this was probably because of habit/habitat-associated parallelisms. Myrothamnaceae were included in Hamamelidales by Cronquist (1981) and Hamamelididae by Takhtajan (1997), probably because of their reduced flowers.
Synonymy: Myrothamnales Reveal - Gunnerineae Shipunov - Myrothamnanae Takhtajan
GUNNERACEAE Meisner, nom. cons. - Gunnera L. - Back to Gunnerales
Perennial (annual) herbs, rhizomatous or stoloniferous; N-fixing Nostoc living intracellularly, infection glands with multicellular papillae; root stele triarch to polyarch, cork cambium 0; vascular cambium 0 [?all taxa], ?cork cambium; vessel elements with simple or few-barred scalariform perforation plates [stems] or scalariform, bars to ca 150 [stolons]; axis polystelic [erect stem] or vascular cylinder [stolons]; nodes multilacunar; stem endodermis +, also low glandular areas; petiole anatomy complex, (endodermis common, in lamina occasional); (lamina amphistomatal); cataphylls common; stolons with opposite scales; leaves spiral, lamina base (cordate), (peltate), colleters +; inflorescence usu. branched-racemose, (plant polygamous), bracteoles 0 (+); K 2 (3), valvate, C 0-2; staminate flowers: (P 4, 0); A 1-2; pollen grains semitectate-reticulate; pistillode +; carpelate flowers: (P 0); staminodes +/(0); G [2], inferior, transverse and alternate with P, uni(bi)locular, stigmas dry; compitum +; ovules 1(-2)/carpel, apical, pendulous, epitropous, micropyle endostomal, outer integument ca 3 cells across, inner integument ?2/ca 4 cells across; embryo sac tetrasporic, 16-celled [Peperomia-type]; fruit drupaceous (nut); seed coat?; endosperm with some starch; n = 17, x = 18 (?9. ?8), nuclear genome [1 C] (0.207-)1.29(-8.031) pg; germination epigeal, phanerocotylar.
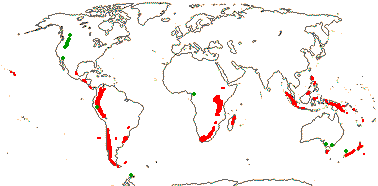
1 [list]/60. Circum S. Pacific, Africa and Madagascar, most New World. Map: see van Balgooy (1975), Trop. Afr. Fl. Pl. Ecol. Distr. vol. 1 (2003) and Wanntorp and Wanntorp (2003); fossil records [green] from the Late Cretaceous onwards, see Jarzen and Dettmann (1989), Dettmann and Jarzen (1990: Fig. 4) and Osborne and Sprent (2002). [Photo - Leaf, Inflorescence]
Age. Fossil pollen of Gunnera is reported in deposits from ca 71 Ma from Namaqualand (Scholtz 1985) and the genus is known from various parts of the South American-Australian area by the later Cretaceous (e.g. Dettmann 1989).
Evolution: Divergence & Distribution. The pollen is distinctive and is known from the Early Cretaceous onwards (Wanntorp et al. 2004b) in all four continents of the Southern Hemisphere, as well as from North America, India and deposits in the Indian (Ninetyeast Ridge) and south Atlantic Oceans (the latter three localities in the Palaeogene - see Jarzen & Dettmann 1989). Wanntorp and Wanntorp (2003) interpreted the distribution of Gunnera in the context of its phylogeny and assuming Gondwanan-age vicariance events (c.f. Beaulieu et al. 2013), so not providing independent evidence for vicariance; see also Sklenár et al. (2011) on the high-altitude South American species.
Ecology & Physiology. Osborne and Sprent (2002) discuss the ecology of the interrelationship between the cyanobacterium and plant.
Plant-Bacterial/Fungal Associations. All taxa have a facultative association with the cyanobacterium, Nostoc (Bergman et al. 1992; Santi et al. 2013: literature; Warshan et al. 2018: relationships of N-fixing members of Nostoc). Glands on the stem immediately below the leaves consist of multicellular papillae; mucilage is secreted, and the epidermis is sloughed off. Nostoc, a filamentous alga, enters the plant via these glands, initially as motile hormogonia, filaments that glide over the surface (Bergman et al. 1992). Johansson and Bergman (1994, and references) and in particular Khamar et al. (2010) and Parniske (2018) describe the establishment of this association; there are different sugars in the mucilage that attracts Nostoc and in the gland tissue in which Nostoc grows (Khamar et al. 2010). It enters plant cells where it loses its motility and produces numerous heterocysts, cells modified to produce an anaerobic environment and in which N fixation occurs; Nostoc may not photosynthesise at all in this stage (Bergman 2002). Söderbäck and Bergman (1993) detail the physiology of the two partners; see also Adams et al. (2006).
Chemistry, Morphology, etc.. There are stipule-like structures on the stem of many species (but this is not an apomorphy for the family) that are interpreted as being cataphylls by Wanntorp et al. (2003). Since they are at least sometimes opposite they may be prophylls; they range in shape from suboblong and entire to deeply laciniate with linear lobes. The lamina varies from 7 mm to 3 m across, and the teeth have a glandular apex that broadens distally; two higher order veins are also involved. The difference in anatomy between stems and stolons is striking, the latter having a central unmedullated stele, at least in section Panka; the roots are triarch to polyarch (Wilkinson 2000). Although Gunnera herteri, sister to the rest of the genus, has normal stem anatomy, it is an annual and its anatomy is conceivably derived. Rutishauser et al. (2004) suggest that the basic plant construction is sympodial.
The difference in size, etc., between the inner and outer tepals (when present) is such that they are sometimes described as sepals and petals (e.g. Wanntorp & Ronse de Craene 2005; Ronse de Craene & Wanntorp 2006; González & Bello 2009). The rather uncommon perfect flowers then have two median sepals, two lateral petals, two stamens opposite the petals, and two carpels also opposite the petals (Ronse de Craene & Wanntorp 2006; González & Bello 2009), indeed, there is considerable infraspecific variation in floral morphology (González & Bello 2009; Mora-Osejo et al. 2011).
Some information is taken from Wilkinson and Wanntorp (2006) and Mora-Osejo et al. (2013), both general, see also Wilkinson (1998: anatomy), Rutishauser et al. (2004: G. herteri floral morphology and development), Wanntorp et al. (2004a: pollen), and Schnegg (1902) and Warming (1913) for ovules, etc..
Phylogeny. For a phylogeny of Gunnera, see Wanntorp et al. (2001, 2002, 2006: summary); relationships are [G. herteri: annual, South America [G. perpensa: Africa [rest of genus]]].
MYROTHAMNACEAE Niedenzu, nom. cons. - Myrothamnus Welwitsch - Back to Gunnerales
Aromatic-resinous shrubs, resurrection plants; essential oils +, gallotannins, myricetin, dihydro/chalcones +; ?cork; vessel elements with scalariform-reticulate perforations, ; pith star-shaped/tetragonal; ?nodes; petiole bundle arcuate; individual epidermal cells resiniferous, palisade tissue 0; plant glabrous; leaves amphistomatic; leaves opposite basally forming a sheath, lamina vernation plicate, secondary veins palmate-flabellate, stipules 2, small, persisting on the petiolar sheath; spikes bracteate, with terminal flowers; P 0; staminate flowers: A 3-4, or (3-)4(-8) and connate, anthers valvate basally, connective produced; pollen in tetrads, triporate, intectate, with clavate projections themselves papillate; pistillode 0; carpelate flowers: staminodes 0; G 3-4, only basally connate, with 5 vascular bundles and surface oil cells, the odd member abaxial, styluli short, recurved, stigma decurrent, in two crests; compitum 0; ovules many/carpel, micropyle bistomal, parietal tissue ca 3 cells across; embryo sac bisporic [chalazal dyad], eight-celled [Allium-type]; fruit follicular (and septicidal); exotestal cells with somewhat thickened outer walls; endosperm development?; n = 10, x = 6 (?7, ?8).
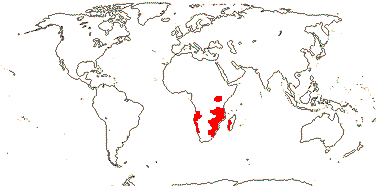
1 [list]/2. Kenya and south tropical Africa, Madagascar. Map: from Puff (1978b) and Trop. Afr. Fl. Pl. Ecol. Distr. vol. 1 (2003). [Photos - Collection]
Evolution: Ecology & Physiology. Myrothamnaceae are homoiochlorophyllous resurrection plants, i.e. the chlorophyll, etc., is not broken down as the plant dries, and they tolerate extreme dessication without any apparent damage. Myrothamnus is the only woody resurrection plant known, in apparently comparable monocots with "trunks" the trunks are made up of dead, persistent leaf bases with roots running through them. The leaves may appear to dry out completely, but they quickly regain turgor, etc., when conditions improve. J. P. Moore et al. (2007) discussed the physiology of Myrothamnus flabellifolia, which shows surprising infraspecific variation. Bianchi et al. (1993) found much glucopyranosyl-ß-glycerol and some trehalose along with the more normal high concentrations of sucrose - stabilizers of phospholipid bilayers, for example - in the dry leaf (see also Farrant 2000; Gaff & Oliver 2013). Mitochondria and perhaps also chloroplasts are surrounded by membranes when the leaf is dry, and the stacking of the chloroplast grana (offset) is very unusual (Wellburn & Wellburn 1976). For water transport, a complicated story, see Sherwin et al. (1998) and in particular Schneider et al. (2003). (Phospho)lipids are much involved in the centrol of water movement and are notably prominent in dried plants, interestingly, water did not flow in the central half of branch xylem, even in well-hydrated plants (Schneider et al. 2003). For Late Embryogenesis Abundant (LEA) genes and gene families, involved in dessication tolerance in angiosperms in general, see Artur et al. (2018).
Chemistry, Morphology, etc.. The stems are narrowly winged. The cork cambium may be superficial, and there are four veins in the leaf sheaths - two bundles go directly to the midrib, and two commissural veins (Grundell 1933), but from transverse sections of older stems there seems to be but a single trace for each leaf (Carlquist 1976b).
Since carpelate flowers lack perianth and staminodes, it is not clear if the ovary is inferior. When flowers are terminal - on the main or lateral axes - there are four "Hochblätter" as well as bracts and bracteoles, and the four carpels are opposite the "Hochblätter" (Jäger-Zürn 1966) which might even be interpreted as perianth or tepals (Wanntorp and Ronse De Craene 2005) and so the ovary would be superior; see also Puff (1978a, 1978b). The carpelate flowers have been described as being zygomorphic (J. P. Moore et al. 2007).
For more information, see Carlquist (1990a: leaf anatomy), also Dahlgren (in Dahlgren & Van Wyk 1988) and Kubitzki (1993b), both general.