EMBRYOPSIDA Pirani & Prado
Gametophyte dominant, independent, multicellular, initially ±globular, not motile, branched; showing gravitropism; glycolate oxidase +, glycolate metabolism in leaf peroxisomes [glyoxysomes], acquisition of phenylalanine lysase* [PAL], flavonoid synthesis*, microbial terpene synthase-like genes +, triterpenoids produced by CYP716 enzymes, CYP73 and phenylpropanoid metabolism [development of phenolic network], xyloglucans in primary cell wall, side chains charged; plant poikilohydrous [protoplasm dessication tolerant], ectohydrous [free water outside plant physiologically important]; thalloid, leafy, with single-celled apical meristem, tissues little differentiated, rhizoids +, unicellular; chloroplasts several per cell, pyrenoids 0; centrioles/centrosomes in vegetative cells 0, microtubules with γ-tubulin along their lengths [?here], interphase microtubules form hoop-like system; metaphase spindle anastral, predictive preprophase band + [with microtubules and F-actin; where new cell wall will form], phragmoplast + [cell wall deposition centrifugal, from around the anaphase spindle], plasmodesmata +; antheridia and archegonia +, jacketed*, surficial; blepharoplast +, centrioles develop de novo, bicentriole pair coaxial, separate at midpoint, centrioles rotate, associated with basal bodies of cilia, multilayered structure + [4 layers: L1, L4, tubules; L2, L3, short vertical lamellae] (0), spline + [tubules from L1 encircling spermatid], basal body 200-250 nm long, associated with amorphous electron-dense material, microtubules in basal end lacking symmetry, stellate array of filaments in transition zone extended, axonemal cap 0 [microtubules disorganized at apex of cilium]; male gametes [spermatozoids] with a left-handed coil, cilia 2, lateral, asymmetrical; oogamy; sporophyte +*, multicellular, growth 3-dimensional*, cuticle +*, plane of first cell division transverse [with respect to long axis of archegonium/embryo sac], sporangium and upper part of seta developing from epibasal cell [towards the archegonial neck, exoscopic], with at least transient apical cell [?level], initially surrounded by and dependent on gametophyte, placental transfer cells +, in both sporophyte and gametophyte, wall ingrowths develop early; suspensor/foot +, cells at foot tip somewhat haustorial; sporangium +, single, terminal, dehiscence longitudinal; meiosis sporic, monoplastidic, MTOC [= MicroTubule Organizing Centre] associated with plastid, sporocytes 4-lobed, cytokinesis simultaneous, preceding nuclear division, quadripolar microtubule system +; wall development both centripetal and centrifugal, 1000 spores/sporangium, sporopollenin in the spore wall* laid down in association with trilamellar layers [white-line centred lamellae; tripartite lamellae]; plastid transmission maternal; nuclear genome [1C] <1.4 pg, main telomere sequence motif TTTAGGG, KNOX1 and KNOX2 [duplication] and LEAFY genes present, ethylene involved in cell elongation; chloroplast genome with close association between trnLUAA and trnFGAA genes [precursors for starch synthesis], tufA, minD, minE genes moved to nucleus; mitochondrial trnS(gcu) and trnN(guu) genes +.
Many of the bolded characters in the characterization above are apomorphies of more or less inclusive clades of streptophytes along the lineage leading to the embryophytes, not apomorphies of crown-group embryophytes per se.
All groups below are crown groups, nearly all are extant. Characters mentioned are those of the immediate common ancestor of the group, [] contains explanatory material, () features common in clade, exact status unclear.
POLYSPORANGIOPHYTA†
Sporophyte well developed, branched, branching dichotomous, potentially indeterminate; hydroids +; stomata on stem; sporangia several, terminal; spore walls not multilamellate [?here].
II. TRACHEOPHYTA / VASCULAR PLANTS
Sporophyte long lived, cells polyplastidic, photosynthetic red light response, stomata open in response to blue light; plant homoiohydrous [water content of protoplasm relatively stable]; control of leaf hydration passive; plant endohydrous [physiologically important free water inside plant]; PIN[auxin efflux facilitators]-mediated polar auxin transport; (condensed or nonhydrolyzable tannins/proanthocyanidins +); borate cross-linked rhamnogalactan II, xyloglucans with side chains uncharged [?level], in secondary walls of vascular and mechanical tissue; lignins +; roots +, often ≤1 mm across, root hairs and root cap +; stem apex multicellular [several apical initials, no tunica], with cytohistochemical zonation, plasmodesmata formation based on cell lineage; vascular development acropetal, tracheids +, in both protoxylem and metaxylem, G- and S-types; sieve cells + [nucleus degenerating]; endodermis +; stomata numerous, involved in gas exchange; leaves +, vascularized, spirally arranged, blades with mean venation density ca 1.8 mm/mm2 [to 5 mm/mm2], all epidermal cells with chloroplasts; sporangia in strobili, sporangia adaxial, columella 0; tapetum glandular; sporophyte-gametophyte junction lacking dead gametophytic cells, mucilage, ?position of transfer cells; MTOCs not associated with plastids, basal body 350-550 nm long, stellate array in transition region initially joining microtubule triplets; archegonia embedded/sunken [only neck protruding]; embryo suspensor +, shoot apex developing away from micropyle/archegonial neck [from hypobasal cell, endoscopic], root lateral with respect to the longitudinal axis of the embryo [plant homorhizic].
[MONILOPHYTA + LIGNOPHYTA]Sporophyte growth ± monopodial, branching spiral; roots endomycorrhizal [with Glomeromycota], lateral roots +, endogenous; G-type tracheids +, with scalariform-bordered pits; leaves with apical/marginal growth, venation development basipetal, growth determinate; sporangium dehiscence by a single longitudinal slit; cells polyplastidic, MTOCs diffuse, perinuclear, migratory; blepharoplasts +, paired, with electron-dense material, centrioles on periphery, male gametes multiciliate; nuclear genome [1C] 7.6-10 pg [mode]; chloroplast long single copy ca 30kb inversion [from psbM to ycf2]; mitochondrion with loss of 4 genes, absence of numerous group II introns; LITTLE ZIPPER proteins.
LIGNOPHYTA†
Sporophyte woody; stem branching axillary, buds exogenous; lateral root origin from the pericycle; cork cambium + [producing cork abaxially], vascular cambium bifacial [producing phloem abaxially and xylem adaxially].
SEED PLANTS† / SPERMATOPHYTA†
Growth of plant bipolar [plumule/stem and radicle/root independent, roots positively geotropic]; plants heterosporous; megasporangium surrounded by cupule [i.e. = unitegmic ovule, cupule = integument]; pollen lands on ovule; megaspore germination endosporic, female gametophyte initially retained on the plant, free-nuclear/syncytial to start with, walls then coming to surround the individual nuclei, process proceeding centripetally.
EXTANT SEED PLANTS
Plant evergreen; nicotinic acid metabolised to trigonelline, (cyanogenesis via tyrosine pathway); microbial terpene synthase-like genes 0; primary cell walls rich in xyloglucans and/or glucomannans, 25-30% pectin [Type I walls]; lignin chains started by monolignol dimerization [resinols common], particularly with guaiacyl and p-hydroxyphenyl [G + H] units [sinapyl units uncommon, no Maüle reaction]; roots often ≥1 mm across, stele diarch to pentarch, xylem and phloem originating on alternating radii, cork cambium deep seated, gravitropism response fast; stem apical meristem complex [with quiescent centre, etc.], plasmodesma density in SAM 1.6-6.2[mean]/μm2 [interface-specific plasmodesmatal network]; eustele +, protoxylem endarch, endodermis 0; wood homoxylous, tracheids and rays alone, tracheid/tracheid pits circular, bordered; mature sieve tube/cell lacking functioning nucleus, sieve tube plastids with starch grains; phloem fibres +; cork cambium superficial; leaf nodes 1:1, a single trace leaving the vascular sympodium; leaf vascular bundles amphicribral; guard cells the only epidermal cells with chloroplasts, stomatal pore with active opening in response to leaf hydration, control by abscisic acid, metabolic regulation of water use efficiency, etc.; branching by axillary buds, exogenous; prophylls two, lateral; leaves with petiole and lamina, development basipetal, lamina simple; sporangia borne on sporophylls; spores not dormant; microsporophylls aggregated in indeterminate cones/strobili; grains monosulcate, aperture in ana- position [distal], primexine + [involved in exine pattern formation with deposition of sporopollenin from tapetum there], exine and intine homogeneous, exine alveolar/honeycomb; ovules with parietal tissue [= crassinucellate], megaspore tetrad linear, functional megaspore single, chalazal, sporopollenin 0; gametophyte ± wholly dependent on sporophyte, development initially endosporic [apical cell 0, rhizoids 0, etc.]; male gametophyte with tube developing from distal end of grain, male gametes two, developing after pollination, with cell walls; embryo cellular ab initio, suspensor short-minute, embryonic axis straight [shoot and root at opposite ends], primary root/radicle produces taproot [= allorhizic], cotyledons 2; embryo ± dormant; chloroplast ycf2 gene in inverted repeat, trans splicing of five mitochondrial group II introns, rpl6 gene absent; ??whole nuclear genome duplication [ζ/zeta duplication event], 2C genome size (0.71-)1.99(-5.49) pg, two copies of LEAFY gene, PHY gene duplications [three - [BP [A/N + C/O]] - copies], 5.8S and 5S rDNA in separate clusters.
IID. ANGIOSPERMAE / MAGNOLIOPHYTA
Lignans, O-methyl flavonols, dihydroflavonols, triterpenoid oleanane, apigenin and/or luteolin scattered, [cyanogenesis in ANA grade?], lignin also with syringyl units common [G + S lignin, positive Maüle reaction - syringyl:guaiacyl ratio more than 2-2.5:1], hemicelluloses as xyloglucans; root cap meristem closed (open); pith relatively inconspicuous, lateral roots initiated immediately to the side of [when diarch] or opposite xylem poles; epidermis probably originating from inner layer of root cap, trichoblasts [differentiated root hair-forming cells] 0, hypodermis suberised and with Casparian strip [= exodermis]; shoot apex with tunica-corpus construction, tunica 2-layered; starch grains simple; primary cell wall mostly with pectic polysaccharides, poor in mannans; tracheid:tracheid [end wall] plates with scalariform pitting, multiseriate rays +, wood parenchyma +; sieve tubes enucleate, sieve plates with pores (0.1-)0.5-10< µm across, cytoplasm with P-proteins, not occluding pores of plate, companion cell and sieve tube from same mother cell; ?phloem loading/sugar transport; nodes 1:?; dark reversal Pfr → Pr; protoplasm dessication tolerant [plant poikilohydric]; stomata randomly oriented, brachyparacytic [ends of subsidiary cells ± level with ends of guard cells], outer stomatal ledges producing vestibule, reduction in stomatal conductance with increasing CO2 concentration; lamina formed from the primordial leaf apex, margins toothed, development of venation acropetal, overall growth ± diffuse, secondary veins pinnate, fine venation hierarchical-reticulate, (1.7-)4.1(-5.7) mm/mm2, vein endings free; flowers perfect, pedicellate, ± haplomorphic, protogynous; parts free, numbers variable, development centripetal; P = T, petal-like, each with a single trace, outer members not sharply differentiated from the others, not enclosing the floral bud; A many, filament not sharply distinguished from anther, stout, broad, with a single trace, anther introrse, tetrasporangiate, sporangia in two groups of two [dithecal], each theca dehiscing longitudinally by a common slit, ± embedded in the filament, walls with at least outer secondary parietal cells dividing, endothecium +, cells elongated at right angles to long axis of anther; tapetal cells binucleate; microspore mother cells in a block, microsporogenesis successive, walls developing by centripetal furrowing; pollen subspherical, tectum continuous or microperforate, ektexine columellate, endexine restricted to the apertural regions, thin, compact, intine in apertural areas thick, orbicules +, pollenkitt +; nectary 0; carpels present, superior, free, several, spiral, ascidiate [postgenital occlusion by secretion], stylulus at most short [shorter than ovary], hollow, cavity not lined by distinct epidermal layer, stigma ± decurrent, carinal, dry; suprastylar extragynoecial compitum +; ovules few [?1]/carpel, marginal, anatropous, bitegmic, micropyle endostomal, outer integument 2-3 cells across, often largely subdermal in origin, inner integument 2-3 cells across, often dermal in origin, parietal tissue 1-3 cells across, nucellar cap?; megasporocyte single, hypodermal, functional megaspore lacking cuticle; female gametophyte lacking chlorophyll, four-celled [one module, egg and polar nuclei sisters]; ovule not increasing in size between pollination and fertilization; pollen grains bicellular at dispersal, germinating in less than 3 hours, siphonogamy, pollen tube unbranched, growing towards the ovule, between cells, growth rate (ca 10-)80-20,000 µm h-1, tube apex of pectins, wall with callose, lumen with callose plugs, penetration of ovules via micropyle [porogamous], whole process takes ca 18 hours, distance to first ovule 1.1-2.1 mm; male gametophytes tricellular, gametes 2, lacking cell walls, ciliae 0, double fertilization +, ovules aborting unless fertilized; fruit indehiscent, P deciduous; mature seed much larger than fertilized ovule, small [<5 mm long], dry [no sarcotesta], exotestal; endosperm +, ?diploid [one polar nucleus + male gamete], cellular, development heteropolar [first division oblique, micropylar end initially with a single large cell, divisions uniseriate, chalazal cell smaller, divisions in several planes], copious, oily and/or proteinaceous, embryo short [<¼ length of seed]; plastid and mitochondrial transmission maternal; Arabidopsis-type telomeres [(TTTAGGG)n]; nuclear genome [2C] (0.57-)1.45(-3.71) [1 pg = 109 base pairs], ??whole nuclear genome duplication [ε/epsilon event]; ndhB gene 21 codons enlarged at the 5' end, single copy of LEAFY and RPB2 gene, knox genes extensively duplicated [A1-A4], AP1/FUL gene, palaeo AP3 and PI genes [paralogous B-class genes] +, with "DEAER" motif, SEP3/LOFSEP and three copies of the PHY gene, [PHYB [PHYA + PHYC]]; chloroplast IR expansions, chlB, -L, -N, trnP-GGG genes 0.
[NYMPHAEALES [AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]]]: wood fibres +; axial parenchyma diffuse or diffuse-in-aggregates; pollen monosulcate [anasulcate], tectum reticulate-perforate [here?]; ?genome duplication; "DEAER" motif in AP3 and PI genes lost, gaps in these genes.
[AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]]: phloem loading passive, via symplast, plasmodesmata numerous; vessel elements with scalariform perforation plates in primary xylem; essential oils in specialized cells [lamina and P ± pellucid-punctate]; tension wood + [reaction wood: with gelatinous fibres, G-fibres, on adaxial side of branch/stem junction]; anther wall with outer secondary parietal cell layer dividing; tectum reticulate; nucellar cap + [character lost where in eudicots?]; 12BP [4 amino acids] deletion in P1 gene.
[MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]] / MESANGIOSPERMAE: benzylisoquinoline alkaloids +; sesquiterpene synthase subfamily a [TPS-a] [?level], polyacetate derived anthraquinones + [?level]; outer epidermal walls of root elongation zone with cellulose fibrils oriented transverse to root axis; P more or less whorled, 3-merous [?here]; pollen tube growth intra-gynoecial; extragynoecial compitum 0; carpels plicate [?here]; embryo sac monosporic [spore chalazal], 8-celled, bipolar [Polygonum type], antipodal cells persisting; endosperm triploid.
[CERATOPHYLLALES + EUDICOTS]: ethereal oils 0 [or next node up]; fruit dry [very labile].
EUDICOTS: (Myricetin +), asarone 0 [unknown in some groups, + in some asterids]; root epidermis derived from root cap [?Buxaceae, etc.]; (vessel elements with simple perforation plates in primary xylem); nodes 3:3; stomata anomocytic; flowers (dimerous), cyclic; protandry common; K/outer P members with three traces, ("C" +, with a single trace); A ?, filaments fairly slender, anthers basifixed; microsporogenesis simultaneous, pollen tricolpate, apertures in pairs at six points of the young tetrad [Fischer's rule], cleavage centripetal, wall with endexine; G with complete postgenital fusion, stylulus/style solid [?here], short [<2 x length of ovary]; seed coat?; palaeotetraploidy event.
[PROTEALES [TROCHODENDRALES [BUXALES + CORE EUDICOTS]]]: (axial/receptacular nectary +).
[TROCHODENDRALES + BUXALES] [CORE EUDICOTS]]: benzylisoquinoline alkaloids 0; euAP3 + TM6 genes [duplication of paleoAP3 gene: B class], mitochondrial rps2 gene lost.
[BUXALES + CORE EUDICOTS]: mitochondrial rps11 gene lost.
CORE EUDICOTS / GUNNERIDAE + DILLENIALES: (ellagic and gallic acids +); leaf margins serrate; compitum + [one position]; micropyle?; γ genome duplication [allopolyploidy, 4x x 2x], x = 3 x 7 = 21, 2C genome size (0.79-)1.05(-1.41) pg, PI-dB motif +; small deletion in the 18S ribosomal DNA common.
[ROSIDS ET AL. + ASTERIDS ET AL.] / PENTAPETALAE / [SANTALALES, CARYOPHYLLALES, SAXIFRAGALES, DILLENIALES, VITALES, ROSIDAE, [BERBERIDOPSIDALES + ASTERIDAE]: root apical meristem closed; (cyanogenesis also via [iso]leucine, valine and phenylalanine pathways); flowers rather stereotyped: 5-merous, parts whorled; P = K + C, K enclosing the flower in bud, with three or more traces, odd K adaxial, C with single trace; A = 2x K/C, in two whorls, alternating, (many, but then usually fasciculate and/or centrifugal); pollen tricolporate; G [(3, 4) 5], when 5 opposite K, whorled, placentation axile, style +, stigma not decurrent, compitum + [one position]; endosperm nuclear/coenocytic; fruit dry, dehiscent, loculicidal [when a capsule]; floral nectaries with CRABSCLAW expression, RNase-based gametophytic incompatibility system present.
Note: In all node characterizations, boldface denotes a possible apomorphy, (....) denotes a feature the exact status of which in the clade is uncertain, [....] includes explanatory material; other text lists features found pretty much throughout the clade. However, the precise node to which many characters, particularly the more cryptic ones, should be assigned is unclear. This is partly because homoplasy is very common, in addition, basic information for all too many characters is very incomplete, frequently coming from taxa well embedded in the clade of interest and so making the position of any putative apomorphy uncertain. Then there are the not-so-trivial issues of how character states are delimited and ancestral states are reconstructed (see above).
Age. The age of this clade has been estimated at some 119.2 Ma (Tank et al. 2015: Table S1, superr + superast; Dilleniaceae 118.1 Ma), or 113 Ma (Leebens-Mack et al. 2005, but sampling); Anderson et al. (2005) suggest a similar figure (stem group to 116 Ma old, crown group diversification by ca 109 My); Chaw et al. (2004: 61 chloroplast genes, sampling poor) date the crown group to 115-110 Ma, and Magallón and Castillo (2009) to around 114.5 Ma. Moore et al. (2010: 95% highest posterior density, see also N. Zhang et al. 2012) suggest crown-group ages of (113-)109(-104) Ma, Bell et al. (2010) ages of (124-)121, 117(-97) Ma, and Magallón et al. (2013, 2015: note topology) ages of (115.9-)110.5-109(-103) Ma and ca 123.7 Ma respectively. Xue et al. (2012: Dilleniaceae not included) suggest the youngest age, some 104.7-101.6 Ma, at ca 166 Ma, the estimate in Z. Wu et al. (2014) is the oldest, while at ca 130 Ma, Foster et al. (2016a: q.v. for details, no Dilleniales) and 128.1-103.5 Ma (Zeng et al. 2017, but c.f. Foster & Ho 2017) are somewhere in between; see also Lutzoni et al. (2018) with an estimate of (125.8-)118.7, 115.3(-109.7) Ma and C. Zhang et al. (2020) with one of ca 125.6 Ma, while Hernández-Hernández et al. (2006) suggest an age of about 92.3 Ma.
Soltis et al. (2008: a variety of estimates) suggested that rosids and asterids in particular diverged 128-114(-84) Ma, and similar ages of (131-)120, 117(-112) and (132-)125, 101(-97) Ma are proposed by Bell et al. (2010). Wikström et al. (2001) suggested ages of (130-)125, 114(-109) Ma, while (128-)127(-124) Ma is the estimate in Wikström et al. (2015), (116.2-)114(-111.3) Ma in Strijk et al. (2019), and ca 89 Ma in the Potato Genome Sequencing Consortium (2011; potato/grape), although the last three in particular did not include Dillenia.
The oldest known core eudicot macrofossil is the well-known Rose Creek fossil from the Cretaceous-Cenomanian some 96-94 Ma (Basinger & Dilcher 1984), quite recently described as Dakotanthus cordiformis by Manchester et al. (2018a). The flower is relatively large compared to the tiny flowers so common in early Cretaceous angiosperms, the five stamens are opposite the petals and there is a well developed nectary, the earliest recorded in the fossil record (Friis et al. 2011). However, with the reinterpretation of flowers attributable to this taxon, the plant appears to have had 10 stamens (the androecium is shown as being obdiplostemonous), fruits are associated with it, and it has been described as Dakotanthus cordiformis (Manchester et al. 2018a). Caliciflora mauldinensis, of about the same age and from Maryland, it also pentapetalan, has a rather different floral morphology, not to mention stellate hairs and a valvate-recurved calyx (Friis et al. 2016). Tropidogyne is an eudicot from Burmese amber and also of about the same age (Chambers et al. 2010; see also Poinar and Chambers 2017), but if Cunoniaceae, its age conflict with other estimates for that clade; Dakotanthus is thought to show a particularly close association (among extant angiosperms) with the southern South American Quillajaceae (Manchester et al. 2018a).
Evolution: Divergence & Distribution. The positions of possible apomorphies/key innovations around here are particularly unclear. Not only is information for all too many critical taxa incomplete, there is also some uncertainty about some relationships in the tree between Ranunculales and Gunnerales, i.e. Buxales and Trochodendrales, and the position of Dilleniales in particular is also unclear, a problem emphasized by studies that focus of relationships along the spine of Pentapetalae (see "Phylogeny" below). However, Sauquet et al. (2017: Supplementary Fig. 5) reconstructed an ancestral floral morphology for this node, although they had some trouble working out what might be going on with the androecium. This floral morphology was also that of several other major nodes, including ... Doyle (2013) suggested that the distinctive pentapetalous flower descibed above evolved from a wind-pollinated, probably dimerous morphology.
Tank et al. (2015) suggest that an increase in the net diversification rate associated with the γ genome duplication event (see Gunneridae) could be linked with the [Superosidae + Superasteridae] node; this increase is placed here, pending clarification of the position of Dilleniales. Indeed, J. W. Clark and Donoghue (2018, q.v. for references) marshal extensive suggestive evidence for diversification and the evolution of extensive morphological disparity around here - for instance, the core eudicots occupy a much greater area of morphospace than the basal eudicots (ibid. Fig. 3). Of course, since clade formation seems to have proceeded apace, genome changes that have affected diversity may have happened independemtly in these clades (Clark & Donoghue 2018). Relocated γ duplicate genes may be important here. They showed relaxed purifying selection, hence there may have been enhancement of amino acid changes and the evolution of novel functions, perhaps contributing to the subsequent diversification of Pentapetalae; relocated γ duplicate genes are significantly enriched in genes with essential functions compared to other γ duplicate genes (Y. Wang et al. 2016).
The flowers of Pentapetalae are very distinctive, as indicated by the characterisation above. Five-merous flowers (K5 C5 A10 is a common combination) preponderate (hence the name "pentapetalae"), but they are uncommon in more basal clades (González & Mello 2009). Compared to many more basal eudicots and monocots, the two perianth whorls are distinctive in that members of each encircle the floral axis, all members of the androecial whorls being adaxial/interior to the petals/inner whorl. Sepals (and bracts) usually have three traces from three gaps and petals have just one (von Balthazar & Endress 2002; Ronse de Craene 2007), however, in several more basal eudicot clades the outer perianth members/sepals have only a single gap (see also Ranunculales); only cases where the vasculature differs are mentioned below. Three-trace petals are sometimes to be found in other eudicots, magnoliids, etc.. There has indeed been much discussion as to the distinction between and evolution of sepals and petals. Petals here may generally be derived from tepals, perhaps ultimately from bracts, not from stamens - possible exceptions include Caryophyllaceae, etc. (Ronse de Craene 2007, 2008, but c.f. Wei & Ronse De Craene 2019).
Taxa that have flowers with many stamens are scattered throughout the core eudicots. The stamens usually develop on common primordia, whether a ring primordium or five or ten separate primordia, when five, the primordia are often opposite the petals, rather than alternating with them, numerous individual stamens developing from these few initial primordia, and development is often centrifugal (secondary or complex polyandry). At maturity, the stamens of Pentapetalae may be more or less connate or in fascicles. This is unlike androecial development in magnoliids and the ANA grade, many Ranunculales, etc., where the numerous stamens develop acropetally from separate primordia, and this is known as primary polyandry (see Corner 1946b; Weberling 1989; Ronse Decraene & Smets 1992b, 1998c; Leins 2000; Endress 2013 and references; Prenner et al. 2008; Rudall 2010: from a morphogenetic point of view). Closely related multistaminate taxa can differ in details of androecial development (e.g. Hufford 1990; Ge et al. 2007). Note that polyandry is decidedly less common in the core asterid/gentianid clade (see discussion there) and it then appears to be linked with increases of numbers of petals and/or carpels (e.g. Nuraliev et al. 2019 and references: Araliaceae), just one of the ways in which polyandry occurs here (Ronse De Craene 2016b).
Tentatively, then, and based largely on observations of gross morphology, there seem to have been major changes in floral organisation at the angiosperm, monocot, commelinid, Pentapetalae, asterid, and the gentianid nodes. As noted under the core eudicots, the floral morphology of extant Gunnerales is very different from that of Pentapetalae and is more similar to that of the eudicots immediately basal to them, so the effects of the genome duplication on floral organization are not apparent there - and the palaeohexaploidy there has been resolved into two events, one of which is at the eudicot node (see above... The presence of a compitum in rosids and the extended clade including asterids might be key innovations for each (Endress 2011a). Few Dilleniaceae have a compitum and its presence is spotty in basal eudicot clades, although Gunneraceae and Sabiaceae, at least, also have a compitum. I have placed the character at this node; strictly speaking Dilleniaceae will then have lost a compitum, although if Dilleniaceae turn out to be sister to other Pentapetalae (although this is now not so likely - see H.-T. Li et al. 2019), it could be placed at the node above or, depending on the optimization, at nodes below. Five-merous flowers and the distinction between sepals and petals are other potential key innovations, whatever the position of Dilleniaceae (Endress 2011a). As we will see below, changes in gross floral morphology may be linked with changes in genes associated with the development of the flower, and these in turn may ultimately go back to the palaeohexaploidy, the gamma triplication event, that has been placed at the Gunnerales node below. Indeed, there may have beeen a shift to the ABCDE floral developmental model, with expression domains fairly discrete, from the fading borders model at the Pentapetalae node (Z. Zhang et al. 2018 and references).
For the development of the flower of Berberidopsis corallina and Aextoxicon (Berberidopsidales), possible "links" in the evolution of the flower of core eudicots, see Ronse De Craene (2004, 2007, 2010). The link is at best thought of as an analogy, since several elements of their floral morphology are probably parallelisms within core eudicots and others may even be reversals; there is considerable variation in floral morphology in this small clade. Unfortunately, relationships around here are unclear. Berberidopsidales may be part of the pectination immediately basal to the asterids, relationships perhaps being [Berberidopsidales [Santalales [Caryophyllales + Asterids]]], however, in H.-T. Li et al. (2019), who are followed below, the positions of the first two orders is reversed (see also W. J. Baker et al. 2021a: Seed Plant Tree), and those relationships are provisionally followed here.
The clarification of the phylogenetic position of Dilleniales, etc. (see also below), is essential for developing a better understanding of floral evolution in this part of the tree. Thus Chase (2005) noted that in Santalales some floral parts, particularly stamens, might have several whorls, and this perhaps suggested that canalisation of floral development there was less than in some other core eudicots; whether Santalales really are different in this respect from other core eudicot groups remains to be established.
Harmomegathic movements of triaperturate pollen grains are discussed by Halbritter and Hesse (2004). For the pollan morphology of a number of the groups just mentioned, see Y. Yu et al. (2018).
Pollination Biology. For the evolution of the RNase-based gametophytic incompatibility system, perhaps a precursor to the sporophytic system (although the two may in fact merely be qualitatively different), see Igic and Kohn (2001: phylogeny of RNases), Steinbachs and Holsinger (2002), Igic et al. (2006), Franklin-Tong and Franklin (2003), Charlesworth et al. (2005) and Hiscock and Tabah (2003).
Genes & Genomes. There has been duplication of all four MADS-box gene classes (A, B, C, E) somewhere around here, and also the floral symmetry genes CYCLOIDEA and DIVARICATA (e.g. Boyden et al. 2010), indeed, this duplication of genes important in determining the identity of the parts of pentapetalous flowers (AP3, AP1, SEP, AG) may well be connected with the γ genome triplication event that is an apomorphy for the core eudicots (Jiao et al. 2012; Vekemans et al. 2012; P. Soltis & Soltis 2016: possible key innovations; J. W. Clark & Donoghue 2018). Thus Hernández-Hernández et al. (2006) and Gloppato and Dornelas (2018) note a duplication in the palaeoAP3/APETALA3-like gene in the ancestor of core eudicots (which the former date as late as ca 92.3 Ma) producing euAP3-like and TM6-like genes, the latter remaining more similar to the palaeoAP3 lineage, and here heterodimerization becomes obligate (see also below. For monosymmetry and the patterns of CYC gene expression, monosymmetric flowers having adaxial/dorsal CYC expression, see Hileman (2014), indeed, a common genetic network involved in floral symmetry seems to occur throughout monocots and eudicots (Madrigal et al. 2019).
Lee et al. (2004) suggest that the CRABS CLAW gene is expressed in core eudicot nectaries (including extrafloral nectaries), or at least in the rosids and asterids that they sampled; it was not expressed in nectaries of Ranunculaceae; what happens in Proteaceae, which has axial nectaries, like rosids (e.g. Smets 1988) and Sabiaceae is unfortunately unknown, moreover, this gene is not expressed in the extrafloral nectaries of Passifloraceae (Krosnick et al. 2008a). Smets et al. (2003) characterise the rosids as having receptacular nectaries, although Vitales have gynoecial nectaries; Proteaceae, and perhaps Sabiaceae, that are basal to rosids also have receptacular nectaries (Smets 1988). (Sucrose synthesis and secretion is similar in the nectaries of Brassicaceae and Solanaceae, extrastaminal and gynoecial nectaries respectively - Lin et al. 2014.) The CRABS CLAW and other genes, including those involved in the RNase-based gametophytic incompatibility system (see above), may have an unambiguous position on the tree, but for now I have tentatively placed only the CRABSCLAW gene in the hierarchy - and wherever it is to be placed, it must have been subsequently lost many times (for nectaries, see also Tölke et al. 2019). In general, to place these genes on the tree and to understand these events, denser sampling is much needed!
Only ca 1/5 as many gene families expanded at this node compared to the eudicot node (110 versus 551), while twice the number contracted (P.-L. Liu et al. 2020: [Vitis + the rest] is the node actually involved).
Chemistry, Morphology, etc.. The distribution of ellagic acid is similar to that of common primordium-type polyandry in the eudicots. Sampling of variation in the root apical meristem is poor, and possible reversals (for which, see Groot et al. 2004) have not been placed on the tree. Scattered in core eudicots, including Arabidopsis, is a particular kind of root hair development in which hairs form above the radial walls of two adjacent underlying cortical cells, the H position, but if there is no underlying radial cortical cell wall, the N position, then no hair develops (Schiefelbein et al. 1997: Arabidopsis; Clowes 2000; Pemberton et al. 2001; Dolan & Costa 2001; C. M. Kim & Dolan 2011 and references). Other core eudicots lack differentiated trichoblasts, root hairs not being so restricted in where they develop.
Phylogeny. Ahhh. Pentapetalae are a strongly supported clade, e.g. Chase et al. (1993), D. Soltis et al. (1997, 1999, 2003a), Hoot et al. (1998), Nandi et al. (1998), and just about all subsequent studies; support was rather weaker in Zhu et al. (2007). Within this large clade, although the rosid and asterid clades are well supported (the former perhaps less so), other relationships were less clear, and they were shown as a hexatomy (there is a nonatomy in Magallón & Sanderson 2001) that involved Crossosomatales, Berberidopsidales, Caryophyllales, Santalales, rosids and asterids in the sixth and earlier versions of this site; Dilleniales and Saxifragales were also of uncertain positions.
Hilu et al. (2003: matK analysis, Schumacheria, the one Dilleniaceae included, was firmly associated with Ericales...) found a possible set of relationships [Rosids [[Dilleniacaeae + Vitaceae] [Saxifragales [Santalales [Berberidopsidales [Caryophyllales + Asterids]]]]]]. D. Soltis et al. (2003a: four-gene analysis, only 54% jacknife support) suggested that Berberidopsidales were sister to the rest of the non-rosid core eudicots. Santalales were associated with asterids, while Saxifragales and Vitales linked with [Dilleniales + Caryophyllales], but with still less support (D. Soltis et al. 2003a). In some studies Dilleniaceae were sister to Caryophyllales, but with only very moderate support; D. Soltis et al. (2003a) provided rather stronger (83% jacknife) support for this position (see also Soltis et al. 2007a: 1.0 p.p.). It also seemed possible that [Caryophyllales + Dilleniales] and Santalales formed a clade (D. Soltis et al. 2000); Carlquist (2006) suggested that non-bordered perforation plates was a possible similarity between Santalales and Caryophyllales. Caryophyllales were also linked with asterids in a large 18S ribosomal DNA analysis (Soltis et al. 1997), albeit support was only weak.
In studies using whole plastomes (Jansen et al. 2006a, esp. 2006b; Hansen et al. 2007; Cai et al. 2007; Ruhlman et al. 2006; Jansen et al. 2007; Moore et al. 2007; Logacheva et al. 2008; Gitzendanner et al. 2018a, b) support for a [Caryophyllales + asterid] clade was stronger, however, Berberidospidales, Dilleniales, Santalales and Saxifragales were not always included. [Caryophyllales + Santalales] were sister to asterids in some analyses in a study that focussed on the position of Cynomoriaceae and Balanophoraceae (Nickrent et al. 2005: again, see sampling). In a study using the mitochondrial gene matR, Caryophyllales repeated as sister to asterids, but with very little support; in other analyses including a reduced sampling and two chloroplast genes Santalales and Dilleniales were also in this area, but again with little support (Zhu et al. 2007). In the combined morphological and molecular study of Nandi et al. (1998) the position of Caryophyllales was uncertain, but this was perhaps partly because the ovules of Rhabdodendraceae, there sister to all other Caryophyllales, were interpreted as being unitegmic; however, subsequent work suggested that Rhabdodendraceae are not sister to all other Caryophyllales, but rather to core Caryophyllales and immediately associated families (see Caryophyllales).
In any event, it seemed increasingly likely that Caryophyllales, whether or not accompanied by a number of other taxa, were sister to asterids (but c.f. Goloboff et al. 2009). In a two-gene study focussing on early-diverging eudicots, Dilleniales, Berberidopsidales, Santalales and Caryophyllales grouped in a pectinate fashion with the asterids, although support was low (Hilu et al. 2008). Wang et al. (2009: 12-gene plus plastid inverted repeat) in a study of the rosids found that Berberidopsidales was sister to a clade made up of the few Caryophyllales and asterids they included. Moore et al. (2008) in a preliminary analysis of whole-chloroplast genome data, suggested that most members of the basal polytomy of the core eudicots could be placed as a series of pectinate branches immediately basal to the asterids; Moore et al. (2010) suggested the relationships [Santalales [Berberidopsidales [Caryophyllales + Asterids]]], the position of Caryophyllales having the least support (see also Barba-Montoya et al. 2018), while in Moore et al. (2011) the position of the middle two orders was inverted (see also Arakaki et al. 2011; Ruhfel et al. 2014; Zeng et al. 2014: suppl. Fig. 14; Magallón et al. 2015: [Berb. + Cary.], Dilleniales sister to all these). Relationships in Bell et al. (2010) were [Berberidopsidales [Caryophyllales, Santalales, asterids]], and in Soltis et al. (2011: little bootstrap support) they were [Santalales [[Berberidopsidales + Caryophyllales] + asterids]], i.e. their superasteridae. The transcriptome analyses of Wickett et al. (2014) also placed Caryophyllales in this area. All this is consistent with many of the earlier, more tentatively suggested relationships. However, a concatenated analysis of 110 single-copy protein sequences suggested that Beta vulgaris (Caryophyllales) was sister to a [rosid + asterid] clade (Dohm et al. 2013), but there may be methodological (see Xi et al. 2013b) and sampling issues here. Moreover, analyses of mitochondrial and nuclear genes have even placed Caryophyllales within superrosids, although support for this position was underwhelming (Sun et al. 2014). Relationshipa are largely unclear in Stull et al. (2018). A plastome analysis by Gitzendanner et al. 2018a, see also b) recovered the relationships [Santalales [Berberidopsidales [Caryophyllales + Asterids]]] as did the comprehensive plastome analysis of H.-T. Li et al. (2021), and with quite strong support. On the other hand, Y.-L. Qiu et al. (2024) recovered the relationships [[Berberidopsidales + Santalales] [[Dilleniales + Caryophyllales] [Asterids]]].
There has been support for placing Crossosomatales as sister to the core malvid group, i.e. Picramniales, Huerteales, Sapindales, etc. (e.g. Zhu et al. 2007; Soltis et al. 2011; Magallón et al. 2015; M. Sun et al. 2016: support weak; H.-T. Li et al. 2021).
Studies on the duplication of the RPB2 gene and subsequent loss of one of the copies (Oxelman et al. 2004; Luo et al. 2007) suggested that Saxifragales and Rosid I + II clades were linked by the loss of the -I copy, which has also occurred in Santalales and Caryophyllales, but not Vitales, Berberidopsidales, Gunnerales or Dilleniales, which have all lost the RPB2-D copy. Saxifragales and Vitales were found to be successively sister to all eudicots minus Gunnerales (mitochondrial gene only), or to all rosids, but with little support (Zhu et al. 2007). For more discussion on the relationships of Saxifragales and Vitales, see the former in particular.
Saxifragales have sometimes been found to be sister to a [Berberidopsidales ... asterid] clade, although with vanishingly low support, and Vitales sister to a rosid clade, but with scarcely any stronger support (Qiu et al. 2010). What are essentially the reverse positions of these two problematic orders were recovered by Soltis et al. (2015, in N. Zhang et al. 2016). Crossosomatales were in a clade basal to Caryophyllales, while other relationships in the rosid II clade were somewhat scrambled, although again with little support (Qiu et al. 2010). The relationships [[Vitaceae + Saxifragales] [Caryophyllales + rosids]] have also popped up (N. Zhang et al. 2012; M. Sun et al. 2104: mitochondrial and nuclear data) as have the relationships [Vitales [rosids + asterids]] (Wickett et al. 2014: transcriptome analyses), but this latter may be a sampling problem, since no Saxifragales, Dilleniales, etc., were included. H.-T. Li et al. (2021) had quite strong/strong/moderate/strongish support for the topology [Dilleniales [Saxifragales [Vitales + other rosids]] in their plastome analysis.
Some morphological evidence, including seed coat anatomy, suggests a more specific relationship between Dilleniales and Vitales. Horne (2006) also listed a number of features linking Dilleniaceae and Rhabdodendraceae, then thought to be sister to all other Caryophyllales, some of which could be features of a [Dilleniales + Caryophyllales] clade, and the status of the others depended on an improved resolution of relationships. These features include absence of tension wood; successive cambia present; vessel elements with simple perforation plates; wood with SiO2 bodies; nodes 3 or more:3 or more; leaves spiral; K persistent in fruit. Since Rhabdodendraceae are now placed sister to the core Caryophyllales and immediately associated families (e.g. Drysdale et al. 2007; Brockington et al. 2007, 2015), the significance of these similarities is unclear. Triterpenoids produced here by a subgroup of CYP716 enzymes are known elsewhere only from Ranunculales and magnoliids (Miettinen et al. 2017).
The position of Dilleniales remained uncertain. Bell et al. (2010) placed them sister to Caryophyllales and Soltis et al. (2011) in a broadly similar position as sister to their superasteridae, and with 87% ML bootstrap support; this position was also found by Arakaki et al. (2011), who included a larger sample of caryophyllalean chloroplast genomes. Similarly Qiu et al. (2010) found a weakly supported [Dilleniales + Berberidopsidales] clade sister to an [asterid [Santalales + Caryophyllales]] clade, but also with very weak support. Moore et al. (2008), Sun et al. (2013: crown-group age 116-112 My) and Ruhfel et al. (2014: not amino acid analyses - sister to asterids) suggested that Dilleniales were sister to rosids, although support could be stronger, while Moore et al. (2011) found a weakly supported [Dilleniales [superrosids + superasterids]] clade (see also Z.-D. Chen et al. 2016). Zeng et al. (2017) found Berberidopsidaceae to be somewhere at the base of Pentapetalae, either sister to the superrosids or superasterids; the former included [Santalales [Vitales [Saxifragales + Rosids]]] the latter [[Dilleniales + Caryophyllales] Asterids]] or [Dilleniales [Caryophyllales + Asterids]]. Gitzendanner et al. (2018a) also placed Dilleniaceae as sister to all other Pentapetalae, while Gitzendanner et al. (2018b) put them sister to the superrosids. H.-T. Li et al. (2019) recovered a well-supported set of relationships [[Dilleniales + superrosids] [Santalales [Berberidopsidales [Caryophyllales + Asterids]]]], this topology being based on plastome data alone. However, in an analysis of 18S/26S nuclear ribosomal data the clades [Dilleniales + Celastrales] and [Caryophyllales + Zygophyllales] were recovered, both embedded in the rosids but with vanishingly little support, while there was some support for the position of Santalales as sister to all other Pentapetalae, and little support for Berberidopsidales as sister to the remaining Pentapetalae (Maia et al. 2014).
Clearly, relationships immediately to the base of the asterids and rosids have been swirling around (not to mention what has been happening within rosids and asterids). O.T.P.T.I. (2019: Fig. 4b, see also Fig. 3i) in their one thousand transcriptome analyses realistically represented relationships around here in a summary tree as a pentatomy made up of asterids, Vitales, Saxifragales, Caryophyllales, and core rosids; they mentioned various other relationships they had obtained, one including [[Gunnerales + Dilleniales] [[Caryophyllales [Berberidopsidales + asterids]] [Santalales [Vitales [Saxifragales + rosids]]]]], so possibly the whole lot should be in a nonatomy...
T. Zhao et al. (2020, esp. 2021) carried out microsynteny analyses (= the local conservation of gene order and content) of the genomes of 123 species, 52 families and 31 orders of flowering plants, and these orders just mentioned, or at least those that they included, again were distinctly peripatetic. Zhao et al. (2021) did not include any Celastrales or Oxalidales in their studies, but Malpighiales moved from a position sister to the N-fixing clade in A.P.G. IV (2016) to a position sister to a clade including Sapindales, etc., similar to the position obtained by Xi et al. (2014) using nuclear data (and also obtained in some other earlier analyses). Relationships in the core eudicots were ...[Vitales [[[Santalales + Saxifragales] Caryophyllales] asterids] [Myrtales [Malpighiales [Sapindales [Malvales + Brassicales]]] [N-fixing clade]]. But other toploplogies were obtained, depending on the data and how they were analysed.
The focus of the nucleome analyses of C. Zhang et al. (2020) was on the asterids, but the tritomy they obtained for the other Pentapetalae that they examined is interesting - [[Vitales + Saxifragales], [Santalales [[Brassicales + Malpighiales] [Dilleniales + Caryophyllales]]], [Berberidopsidales + asterids].
Finally, W. J. Baker et al. (2021a) in their preliminary Angiosperms353 analysis found a clade [Gunnerales + Dilleniales] (which had vanishingly little support) sister to a [rosids + asterids] clade. Z. Xu et al. (2024) noted that topologies of core eudicots in analyses based on nucleotide sequences, nucleotide sequences without third codon positions, and amino acid sequences differed, especially the last two.
Conclusion: Things are pretty much up in the air. Relationships in this area in APG IV (2016) are [[Gunnerales [Dilleniales, [Saxifragales [Vitales [[Zygophyllales [[Fabales [Rosales [Fagales + Cucurbitales]]]] [[Celastrales [Malpighiales + Oxalidales]]]] [[Myrtales + Geraniales] [Crossosomatales [Picramniales [Sapindales [Huerteales [Malvales + Brassicales]]]]]]]]], [Berberidopsidales [Santalales [Caryophyllales + Asterids]]]. However, there was talk in 2020 of a major analysis of plastome data, including plastomes from nearly all angiosperm families apart from some holoparasitic/-holomycoheterotrophic taxa. In addition, an extended analysis of Angiosperm353 data, with quite dense infrafamilial sampling, was also in the works. See also G. Zhang and Ma (2024: esp. Figs 3-5) for a summary of nuclear genome-based relationships suggested in the pre 2024 literature.
And the results? H.-T. Li et al. (2021) included plastomes from 4,498 species representing all families apart from those of six mycoheterotrophic families that did not behave and had to be excluded. In the recent Angiosperms353 probe set analysis (Zuntini et al. 2024: 7923 genera, see Extended Fig. 1, Suppl. Fig. 4, etc. - the topologies of APG IV 2016 and Li et al. 2021 are included in Figs. 1 and 4 for comparison) relationships in the general rosid area that were recovered are rather different compared to those in A.P.G. IV. Zuntini et al. (2024) recovered the relationships [[Gunnerales + Dilleniales] [[Caryophyllales [Santalales [Vitales [Saxifragales [[Crossosomatales + Geraniales] [[Myrtales + Zygophyllales](support for the placement of these two very weak) [[[Fabales + Fagales] [Rosales + Cucurbitales]] [[Picramniales [Huerteales [Oxalidales [Sapindales [Malvales + Brassicales]]]] [[Celastrales [Apodanthaceae [Malpighiales + Huaceae (support for the placement of the last very weak)]]]]]]]]]]] [Berberidopsidales + Asterids]]]. Finally, relationships in Y.-L. Qiu et al. (2024) are [Vitales [Myrtales [[Geraniales + Crossosomatales] [Zygophyllales [[Fabales + Rosales] [Fagales + Cucurbitales]]]]]]]] [[Sapindales [Huerteales [Brassicales + Malvales]]] [Celastrales (inc. Lepidobotyraceae) [Oxalidales (inc. Huaceae) + Malpighiales]]]] [[Berberidopsidales + Santalales] [[Dilleniales + Caryophyllales] [Asterids]]], and with different topologies at all levels of the hierarchy. Four orders have been picked out above to help see the differences between the two topologies, but note that some ten of the nodes in Zuntini et al. (2024) just mentioned, around 40% of the total, could be better supported. However, if relationships in Zuntini et al. (2024) are confirmed then a few new angiosperm orders may be needed, not to mention substantial rearrangements in the tree. Altogther, rather mind-numbing. For further discussion about major patterns of relationships within Pentapetalae, see asterids, euasterids and Saxifragales-rosids.
Classification. In Versions 8 and earlier of this site this was called the core eudicot clade, largely because the evolution of the "typical" core eudicot flower can be pegged to this node; the current delimitation of core eudicots refers to the immediately more inclusive clade that is molecularly quite well supported but that is perhaps morphologically less interesting, that is, Gunneridae.
[DILLENIALES [SAXIFRAGALES + ROSIDS]] - if this clade exists: stipules + [usually apparently inserted on the stem].
Evolution: Moore et al. (2010) suggest ages of (112-)108(-103) Ma for the crown group; ca 164 Ma is the estimate in Z. Wu et al. (2014).
DILLENIALES Berchtold & J. Presl - Main Tree.
Just the one family, 11 genera, ca 360 species.
Note: In all node characterizations, boldface denotes a possible apomorphy, (....) denotes a feature the exact status of which in the clade is uncertain, [....] includes explanatory material; other text lists features found pretty much throughout the clade. However, the precise node to which many characters, particularly the more cryptic ones, should be assigned is unclear. This is partly because homoplasy is very common, in addition, basic information for all too many characters is very incomplete, frequently coming from taxa well embedded in the clade of interest and so making the position of any putative apomorphy uncertain. Then there are the not-so-trivial issues of how character states are delimited and ancestral states are reconstructed (see above).
Includes Dilleniaceae.
Synonymy: Dillenianae Takhtajan - Dilleniidae Reveal & Takhtajan
DILLENIACEAE Salisbury - Back to Dilleniales
Trees and shrubs; distinctive flavonols, myricetin, ellagic acid +; primary stem with continuous vascular cylinder; cork cambium deep-seated; vessel elements with simple and scalariform perforation plates; true tracheids +; raphides +, also common in wood; rays often broad; (silica bodies +); ?nodes, petiole bundle annular; epidermis silicified; branching from previous flush; hairs unicellular, silicified; leaves spiral, lamina vernation conduplicate(-plicate), surface often scabrid, margins toothed, teeth with clear glandular expanded apex, secondary veins parallel, proceeding straight to the teeth [percurrent, craspedodromous], tertiary venation ± scalariform, fine veins areolate, teeth with clear glandular expanded apex, base broad, stipules 0; inflorescence often terminal; pedicels articulated, bracteoles 0-2; flowers often yellow, lasting one day; K (3-)5(-20), (with 1 trace), large, C (2-)5, quincuncial, crumpled in bud (not); androecium from a ring primordium or fasciculate, fascicles opposite K, supplied by trunk bundles, A many, often centrifugal, (staminodes +), anthers basifixed, epidermis well developed, tanniniferous; pollen tectum punctate-reticulate; nectary 0; G (1-3)4-8[-20], (when = C, opposite them), (odd member adaxial), compitum 0, styluli long, stigma punctate, wet [1 record]; ovules many/carpel, apotropous, often campylotropous (straight), micropyle zigzag or exostomal, outer integument 2(-3) cells across, inner integument 2-6 cells across, parietal tissue 6-14 cells across, nucellar cap ca 2 cells across, chalazal area massive; (megaspore mother cells several); K ± accrescent; fruit a follicle, aril +, funicular, often laciniate, exotesta often fleshy, endotesta ± palisade, massively lignified, cutinized [?all], exotegmen usu. tracheidal, with spiral/annular thickenings, endotegmen tanniniferous/0; zygote with distinctive wall and protrusions into the endosperm ["mantle"]; x = 8/9/6, nuclear genome [1 C] (0>022-)1.177(-62.35) pg; germination phanerocotylar.
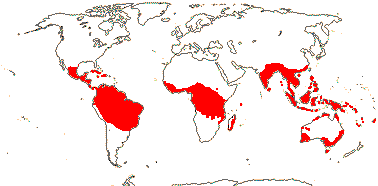
11 [list: to subfamilies]/600 - four groups below. Tropical and warm temperate. Map: from van Steenis and van Balgooy (1966), van Balgooy (1975), Heywood (1978), Trop. Afr. Fl. Pl. Ecol. Distr. vol. 1 (2003) and Horn (2009). [Photos - Collection.]
Age. The stem age may be ca 114.75 Ma and the crown age only ca 52.5 Ma (Magallón & Castillo 2009).
1. Delimoideae Burnett - Tetracera L. —— Synonymy: Delimaceae Martius
Lianes; (successive cambia +); vessel elements usu. with simple perforation plates; nodes 3:3; (petiole bundle strongly arcuate and with adaxial bundles); stomata paracytic; (plant functionally dioecious; pollen in female flowers ± pantoporate, infertile); apex of filament/connective expanded, thecae ± separate; G 1-5(-8), (stigma peltate); endotesta poorly differentiated; n = ?
1/44. Pantropical.
[Doliocarpoideae [Hibbertioideae + Dillenioideae]]: stomata usu. anomocytic;
2. Doliocarpoideae J. W. Horn —— Synonymy: Soramiaceae Martynov
Lianes; (successive cambia +); small vessel elements with scalariform perforation plates; nodes 5:5 (3:3, 7:7); (petiole with medullary bundles); (base surrounding stem, flanges persistent); prophylls 0/reduced [no axillary buds]; (filaments swollen towards apex); G 1-2(-3), (compitum + - Pinzona), stigma peltate-infundibular; ovules 2/carpel, collateral, one epitropous, the other apotropous; (2 K much accrescent, thin, surrounding fruit; dehiscence irregular - Davilla); n = 13.
5/75: Doliocarpus (40), Davilla (28). Neotropical.
[Hibbertioideae + Dillenioideae]: vessel elements usu. with scalariform perforation plates; (flowers monosymmetric, A opposite the median-abaxial C); (A fasciculate), anther dehiscence ± porose); pollen grains colpate.
3. Hibbertioideae J. W. Horn - Hibbertia Andrews —— Synonymy: Hibbertiaceae J. Agardh
Trees to (rhizomatous) shrubs or subshrubs (lianes), (leaves reduced, stems photosynthetic); nodes 3:3, 1:1; petiole bundles various; (hairs hooked/± stellate/fasciculate); lamina (margin entire), (secondary veins descending), tertiary venation not scalariform, (areoles weakly developed), leaf base not sheathing; A (monosymmetric), (1-5)-200+, (obdiplostemonous), (outer/inner staminodes +), (outer A basally connate), (filaments swollen); G (1-)2-5(-15); (1-) ovules/carpel; endosperm starchy; n = 4-6, 8-10, 12, 13, nuclear genome [1 C] ca 2.64 pg [H. scandens].
1/375 (+ >100 undescribed). Madagascar to New Guinea, New Caledonia (25) and Fiji, but nearly all endemic to Australia.
4. Dillenioideae Burnett
Trees, shrubs, (herbaceous, rhizomatous - Acrotrema); cork cambium superficial; nodes 5<:5< (1:1); (petiole with medullary bundles); leaves (deciduous), (blade pinnate - Acrotrema), (base surrounding stem), (petiolar flanges + deciduous or not); A (monosymmetric), 40-200+; pollen grains tricolpate; G (2-)5-15(-many), often ± connate, compitum +, (with a central receptacular cone); (ovule 1/carpel); (calyx massively accrescent and surrounding fruit); exotestal cells (with hairs), (large, tanniniferous, becoming flattened - Dillenia); n = 13, 15, 16, 24, etc.
4/75. Dillenia (60). Madagascar to Fiji, most Indo-Malesian, few Australia.
Evolution: Divergence & Distribution. Species distributions/relationships of Hibbertia within Australia seem to fit the peripheral vicariance pattern, species now being centred on the periphery of the continent after the drying out of the centre, a process that began in the Eocene (Nge et al. 2021c).
Horne (2009) provides phylogenetic optimisations for a number of characters in the family, while Y. Yu et al. (2018) discuss pollen evolution.
Ecology & Physiology. There are about 100 species of stem-twining lianes in the family, most in Delimoideae and Doliocarpoideae. These often have vessels with simple perforation plates, and a correlation between the liane habit and simple vessels has been noted here and elsewhere (Carlquist 1991b; Isnard & Feild 2015). Interestingly, the direction of twining seems not to be fixed in the family (Burnham et al. 2019).
Pollination Biology & Seed Dispersal. Buzz pollination by bees is common in Dilleniaceae, although some species of Hibbertia, for example, have anthers dehiscing longitudinally (Endress 1997b; Tucker & Bernhardt 2000; Horn 2007 and references).
The dispersal unit is generally the arillate seed, which is endozoochorous, eaten by birds or monkeys (e.g. Dillenia), or myrmecochorous, as in Hibbertia in particular (Lengyel et al. 2009, 2010).
Plant-Animal Interactions. Caterpillars of the tortricid Phricanthini moths are known only from Dilleniaceae (Powell et al. 1999).
Chemistry, Morphology, etc.. There are often sclereids in the pith. Hibbertia is very variable vegetatively. Species of Hibbertia sect. Pachynemahave very much reduced leaves and winged, photosynthetic stems; the plants are souped-up inflorescences. Rury and Dickison (1977) describe the diversity of leaf venation patterns in the genus; the leaf teeth may be hydathodal (Rios et al. 2020).
Hibbertia is very variable florally (Hammer et al. 2025). The monosymmetric flowers of Didesmandra aspera have two bundles of stamens on the functionally upper side of the flower, in each there is a single fertile stamen longer than the rest; the flower is drawn as if the plane of symmetry in horizontal (Stapf 1900); the monosymmetric flowers of Schumacheria have only a single staminal bundle in which all stamens are about the same lengths. The androecium is supplied by a whorl of stamen trunks which branch irregulary to supply the individual stamens, and when the androecium is fasciculate, stamens of a single fascicle may be supplied by more than one stamen trunk (C. L. Wilson 1965 [Hibbertia], 1974c [the rest]).
See Dickison (1971b), Kubitzki (1971), Stebbins and Hoogland (1976: Hibbertia), and Horn (2006, 2009) for general information, Dickison (1969) for nodal anatomy, which is variable, Dickison et al. (1978) for vascular anatomy in Hibbertia, Paetow (1931), Endress (1997b) and Tucker and Bernhardt (2000) for floral morphology, Dickison (1970: stamens, etc.), Dickison et al. (1982) and Rakarcha et al. (2018) for pollen, Schnarf (1924), Sastri (1958a) and Swamy and Periasamy (1955) for embryology, etc., and Schatral (1995) for seed anatomy.
Phylogeny. For the relationships above, well supported, I follow Horn (2002, 2009); Hibbertia is paraphyletic, Pachynema being embedded in it, while the status of Acrotrema (Dillenioideae) is unclear. Hammer et al. (2025) looked at relationships in Hibbertia (95 spp.,300 nuclear, 60 plastome sequences from Angiosperms353 and Ozbaits analyses) and found the four subgenera held up, and there were 14other major clades that were on the whole well supported (2 species unplaced), however, relationships between these clades was often unclear and there was substantial nuclear-plastome tree conflict.
Classification. See Horn (2009).