EMBRYOPSIDA Pirani & Prado
Gametophyte dominant, independent, multicellular, initially ±globular, not motile, branched; showing gravitropism; glycolate oxidase +, glycolate metabolism in leaf peroxisomes [glyoxysomes], acquisition of phenylalanine lysase* [PAL], flavonoid synthesis*, microbial terpene synthase-like genes +, triterpenoids produced by CYP716 enzymes, CYP73 and phenylpropanoid metabolism [development of phenolic network], xyloglucans in primary cell wall, side chains charged; plant poikilohydrous [protoplasm dessication tolerant], ectohydrous [free water outside plant physiologically important]; thalloid, leafy, with single-celled apical meristem, tissues little differentiated, rhizoids +, unicellular; chloroplasts several per cell, pyrenoids 0; centrioles/centrosomes in vegetative cells 0, microtubules with γ-tubulin along their lengths [?here], interphase microtubules form hoop-like system; metaphase spindle anastral, predictive preprophase band + [with microtubules and F-actin; where new cell wall will form], phragmoplast + [cell wall deposition centrifugal, from around the anaphase spindle], plasmodesmata +; antheridia and archegonia +, jacketed*, surficial; blepharoplast +, centrioles develop de novo, bicentriole pair coaxial, separate at midpoint, centrioles rotate, associated with basal bodies of cilia, multilayered structure + [4 layers: L1, L4, tubules; L2, L3, short vertical lamellae] (0), spline + [tubules from L1 encircling spermatid], basal body 200-250 nm long, associated with amorphous electron-dense material, microtubules in basal end lacking symmetry, stellate array of filaments in transition zone extended, axonemal cap 0 [microtubules disorganized at apex of cilium]; male gametes [spermatozoids] with a left-handed coil, cilia 2, lateral, asymmetrical; oogamy; sporophyte +*, multicellular, growth 3-dimensional*, cuticle +*, plane of first cell division transverse [with respect to long axis of archegonium/embryo sac], sporangium and upper part of seta developing from epibasal cell [towards the archegonial neck, exoscopic], with at least transient apical cell [?level], initially surrounded by and dependent on gametophyte, placental transfer cells +, in both sporophyte and gametophyte, wall ingrowths develop early; suspensor/foot +, cells at foot tip somewhat haustorial; sporangium +, single, terminal, dehiscence longitudinal; meiosis sporic, monoplastidic, MTOC [= MicroTubule Organizing Centre] associated with plastid, sporocytes 4-lobed, cytokinesis simultaneous, preceding nuclear division, quadripolar microtubule system +; wall development both centripetal and centrifugal, 1000 spores/sporangium, sporopollenin in the spore wall* laid down in association with trilamellar layers [white-line centred lamellae; tripartite lamellae]; plastid transmission maternal; nuclear genome [1C] <1.4 pg, main telomere sequence motif TTTAGGG, KNOX1 and KNOX2 [duplication] and LEAFY genes present, ethylene involved in cell elongation; chloroplast genome with close association between trnLUAA and trnFGAA genes [precursors for starch synthesis], tufA, minD, minE genes moved to nucleus; mitochondrial trnS(gcu) and trnN(guu) genes +.
Many of the bolded characters in the characterization above are apomorphies of more or less inclusive clades of streptophytes along the lineage leading to the embryophytes, not apomorphies of crown-group embryophytes per se.
All groups below are crown groups, nearly all are extant. Characters mentioned are those of the immediate common ancestor of the group, [] contains explanatory material, () features common in clade, exact status unclear.
POLYSPORANGIOPHYTA†
Sporophyte well developed, branched, branching dichotomous, potentially indeterminate; hydroids +; stomata on stem; sporangia several, terminal; spore walls not multilamellate [?here].
II. TRACHEOPHYTA / VASCULAR PLANTS
Sporophyte long lived, cells polyplastidic, photosynthetic red light response, stomata open in response to blue light; plant homoiohydrous [water content of protoplasm relatively stable]; control of leaf hydration passive; plant endohydrous [physiologically important free water inside plant]; PIN[auxin efflux facilitators]-mediated polar auxin transport; (condensed or nonhydrolyzable tannins/proanthocyanidins +); borate cross-linked rhamnogalactan II, xyloglucans with side chains uncharged [?level], in secondary walls of vascular and mechanical tissue; lignins +; roots +, often ≤1 mm across, root hairs and root cap +; stem apex multicellular [several apical initials, no tunica], with cytohistochemical zonation, plasmodesmata formation based on cell lineage; vascular development acropetal, tracheids +, in both protoxylem and metaxylem, G- and S-types; sieve cells + [nucleus degenerating]; endodermis +; stomata numerous, involved in gas exchange; leaves +, vascularized, spirally arranged, blades with mean venation density ca 1.8 mm/mm2 [to 5 mm/mm2], all epidermal cells with chloroplasts; sporangia in strobili, sporangia adaxial, columella 0; tapetum glandular; sporophyte-gametophyte junction lacking dead gametophytic cells, mucilage, ?position of transfer cells; MTOCs not associated with plastids, basal body 350-550 nm long, stellate array in transition region initially joining microtubule triplets; archegonia embedded/sunken [only neck protruding]; embryo suspensor +, shoot apex developing away from micropyle/archegonial neck [from hypobasal cell, endoscopic], root lateral with respect to the longitudinal axis of the embryo [plant homorhizic].
[MONILOPHYTA + LIGNOPHYTA]Sporophyte growth ± monopodial, branching spiral; roots endomycorrhizal [with Glomeromycota], lateral roots +, endogenous; G-type tracheids +, with scalariform-bordered pits; leaves with apical/marginal growth, venation development basipetal, growth determinate; sporangium dehiscence by a single longitudinal slit; cells polyplastidic, MTOCs diffuse, perinuclear, migratory; blepharoplasts +, paired, with electron-dense material, centrioles on periphery, male gametes multiciliate; nuclear genome [1C] 7.6-10 pg [mode]; chloroplast long single copy ca 30kb inversion [from psbM to ycf2]; mitochondrion with loss of 4 genes, absence of numerous group II introns; LITTLE ZIPPER proteins.
LIGNOPHYTA†
Sporophyte woody; stem branching axillary, buds exogenous; lateral root origin from the pericycle; cork cambium + [producing cork abaxially], vascular cambium bifacial [producing phloem abaxially and xylem adaxially].
SEED PLANTS† / SPERMATOPHYTA†
Growth of plant bipolar [plumule/stem and radicle/root independent, roots positively geotropic]; plants heterosporous; megasporangium surrounded by cupule [i.e. = unitegmic ovule, cupule = integument]; pollen lands on ovule; megaspore germination endosporic, female gametophyte initially retained on the plant, free-nuclear/syncytial to start with, walls then coming to surround the individual nuclei, process proceeding centripetally.
EXTANT SEED PLANTS
Plant evergreen; nicotinic acid metabolised to trigonelline, (cyanogenesis via tyrosine pathway); microbial terpene synthase-like genes 0; primary cell walls rich in xyloglucans and/or glucomannans, 25-30% pectin [Type I walls]; lignin chains started by monolignol dimerization [resinols common], particularly with guaiacyl and p-hydroxyphenyl [G + H] units [sinapyl units uncommon, no Maüle reaction]; roots often ≥1 mm across, stele diarch to pentarch, xylem and phloem originating on alternating radii, cork cambium deep seated, gravitropism response fast; stem apical meristem complex [with quiescent centre, etc.], plasmodesma density in SAM 1.6-6.2[mean]/μm2 [interface-specific plasmodesmatal network]; eustele +, protoxylem endarch, endodermis 0; wood homoxylous, tracheids and rays alone, tracheid/tracheid pits circular, bordered; mature sieve tube/cell lacking functioning nucleus, sieve tube plastids with starch grains; phloem fibres +; cork cambium superficial; leaf nodes 1:1, a single trace leaving the vascular sympodium; leaf vascular bundles amphicribral; guard cells the only epidermal cells with chloroplasts, stomatal pore with active opening in response to leaf hydration, control by abscisic acid, metabolic regulation of water use efficiency, etc.; branching by axillary buds, exogenous; prophylls two, lateral; leaves with petiole and lamina, development basipetal, lamina simple; sporangia borne on sporophylls; spores not dormant; microsporophylls aggregated in indeterminate cones/strobili; grains monosulcate, aperture in ana- position [distal], primexine + [involved in exine pattern formation with deposition of sporopollenin from tapetum there], exine and intine homogeneous, exine alveolar/honeycomb; ovules with parietal tissue [= crassinucellate], megaspore tetrad linear, functional megaspore single, chalazal, sporopollenin 0; gametophyte ± wholly dependent on sporophyte, development initially endosporic [apical cell 0, rhizoids 0, etc.]; male gametophyte with tube developing from distal end of grain, male gametes two, developing after pollination, with cell walls; embryo cellular ab initio, suspensor short-minute, embryonic axis straight [shoot and root at opposite ends], primary root/radicle produces taproot [= allorhizic], cotyledons 2; embryo ± dormant; chloroplast ycf2 gene in inverted repeat, trans splicing of five mitochondrial group II introns, rpl6 gene absent; ??whole nuclear genome duplication [ζ/zeta duplication event], 2C genome size (0.71-)1.99(-5.49) pg, two copies of LEAFY gene, PHY gene duplications [three - [BP [A/N + C/O]] - copies], 5.8S and 5S rDNA in separate clusters.
IID. ANGIOSPERMAE / MAGNOLIOPHYTA
Lignans, O-methyl flavonols, dihydroflavonols, triterpenoid oleanane, apigenin and/or luteolin scattered, [cyanogenesis in ANA grade?], lignin also with syringyl units common [G + S lignin, positive Maüle reaction - syringyl:guaiacyl ratio more than 2-2.5:1], hemicelluloses as xyloglucans; root cap meristem closed (open); pith relatively inconspicuous, lateral roots initiated immediately to the side of [when diarch] or opposite xylem poles; epidermis probably originating from inner layer of root cap, trichoblasts [differentiated root hair-forming cells] 0, hypodermis suberised and with Casparian strip [= exodermis]; shoot apex with tunica-corpus construction, tunica 2-layered; starch grains simple; primary cell wall mostly with pectic polysaccharides, poor in mannans; tracheid:tracheid [end wall] plates with scalariform pitting, multiseriate rays +, wood parenchyma +; sieve tubes enucleate, sieve plates with pores (0.1-)0.5-10< µm across, cytoplasm with P-proteins, not occluding pores of plate, companion cell and sieve tube from same mother cell; ?phloem loading/sugar transport; nodes 1:?; dark reversal Pfr → Pr; protoplasm dessication tolerant [plant poikilohydric]; stomata randomly oriented, brachyparacytic [ends of subsidiary cells ± level with ends of guard cells], outer stomatal ledges producing vestibule, reduction in stomatal conductance with increasing CO2 concentration; lamina formed from the primordial leaf apex, margins toothed, development of venation acropetal, overall growth ± diffuse, secondary veins pinnate, fine venation hierarchical-reticulate, (1.7-)4.1(-5.7) mm/mm2, vein endings free; flowers perfect, pedicellate, ± haplomorphic, protogynous; parts free, numbers variable, development centripetal; P = T, petal-like, each with a single trace, outer members not sharply differentiated from the others, not enclosing the floral bud; A many, filament not sharply distinguished from anther, stout, broad, with a single trace, anther introrse, tetrasporangiate, sporangia in two groups of two [dithecal], each theca dehiscing longitudinally by a common slit, ± embedded in the filament, walls with at least outer secondary parietal cells dividing, endothecium +, cells elongated at right angles to long axis of anther; tapetal cells binucleate; microspore mother cells in a block, microsporogenesis successive, walls developing by centripetal furrowing; pollen subspherical, tectum continuous or microperforate, ektexine columellate, endexine restricted to the apertural regions, thin, compact, intine in apertural areas thick, orbicules +, pollenkitt +; nectary 0; carpels present, superior, free, several, spiral, ascidiate [postgenital occlusion by secretion], stylulus at most short [shorter than ovary], hollow, cavity not lined by distinct epidermal layer, stigma ± decurrent, carinal, dry; suprastylar extragynoecial compitum +; ovules few [?1]/carpel, marginal, anatropous, bitegmic, micropyle endostomal, outer integument 2-3 cells across, often largely subdermal in origin, inner integument 2-3 cells across, often dermal in origin, parietal tissue 1-3 cells across, nucellar cap?; megasporocyte single, hypodermal, functional megaspore lacking cuticle; female gametophyte lacking chlorophyll, four-celled [one module, egg and polar nuclei sisters]; ovule not increasing in size between pollination and fertilization; pollen grains bicellular at dispersal, germinating in less than 3 hours, siphonogamy, pollen tube unbranched, growing towards the ovule, between cells, growth rate (ca 10-)80-20,000 µm h-1, tube apex of pectins, wall with callose, lumen with callose plugs, penetration of ovules via micropyle [porogamous], whole process takes ca 18 hours, distance to first ovule 1.1-2.1 mm; male gametophytes tricellular, gametes 2, lacking cell walls, ciliae 0, double fertilization +, ovules aborting unless fertilized; fruit indehiscent, P deciduous; mature seed much larger than fertilized ovule, small [<5 mm long], dry [no sarcotesta], exotestal; endosperm +, ?diploid [one polar nucleus + male gamete], cellular, development heteropolar [first division oblique, micropylar end initially with a single large cell, divisions uniseriate, chalazal cell smaller, divisions in several planes], copious, oily and/or proteinaceous, embryo short [<¼ length of seed]; plastid and mitochondrial transmission maternal; Arabidopsis-type telomeres [(TTTAGGG)n]; nuclear genome [2C] (0.57-)1.45(-3.71) [1 pg = 109 base pairs], ??whole nuclear genome duplication [ε/epsilon event]; ndhB gene 21 codons enlarged at the 5' end, single copy of LEAFY and RPB2 gene, knox genes extensively duplicated [A1-A4], AP1/FUL gene, palaeo AP3 and PI genes [paralogous B-class genes] +, with "DEAER" motif, SEP3/LOFSEP and three copies of the PHY gene, [PHYB [PHYA + PHYC]]; chloroplast IR expansions, chlB, -L, -N, trnP-GGG genes 0.
[NYMPHAEALES [AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]]]: wood fibres +; axial parenchyma diffuse or diffuse-in-aggregates; pollen monosulcate [anasulcate], tectum reticulate-perforate [here?]; ?genome duplication; "DEAER" motif in AP3 and PI genes lost, gaps in these genes.
[AUSTROBAILEYALES [MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]]]: phloem loading passive, via symplast, plasmodesmata numerous; vessel elements with scalariform perforation plates in primary xylem; essential oils in specialized cells [lamina and P ± pellucid-punctate]; tension wood + [reaction wood: with gelatinous fibres, G-fibres, on adaxial side of branch/stem junction]; anther wall with outer secondary parietal cell layer dividing; tectum reticulate; nucellar cap + [character lost where in eudicots?]; 12BP [4 amino acids] deletion in P1 gene.
[MONOCOTS [[CHLORANTHALES + MAGNOLIIDS] [CERATOPHYLLALES + EUDICOTS]]] / MESANGIOSPERMAE: benzylisoquinoline alkaloids +; sesquiterpene synthase subfamily a [TPS-a] [?level], polyacetate derived anthraquinones + [?level]; outer epidermal walls of root elongation zone with cellulose fibrils oriented transverse to root axis; P more or less whorled, 3-merous [?here]; pollen tube growth intra-gynoecial; extragynoecial compitum 0; carpels plicate [?here]; embryo sac monosporic [spore chalazal], 8-celled, bipolar [Polygonum type], antipodal cells persisting; endosperm triploid.
MONOCOTYLEDONS / MONOCOTYLEDONEAE / LILIANAE Takhtajan
Plant herbaceous, perennial, rhizomatous, growth sympodial; non-hydrolyzable tannins [(ent-)epicatechin-4] +, neolignans 0, CYP716 triterpenoid enzymes 0, benzylisoquinoline alkaloids 0, hemicelluloses as xylan, cell wall also with (1->3),(1->4)-ß-D-MLGs [Mixed-Linkage Glucans]; root epidermis developed from outer layer of cortex; endodermal cells with U-shaped thickenings; cork cambium [uncommon] superficial; stele oligo- to polyarch, medullated [with prominent pith], lateral roots arise opposite phloem poles; stem primary thickening meristem +; vascular development bidirectional, bundles scattered, (amphivasal), vascular cambium 0 [bundles closed]; tension wood 0; vessel elements in roots with scalariform and/or simple perforations; tracheids only in stems and leaves; sieve tube plastids with cuneate protein crystals alone; ?nodal anatomy; stomata oriented parallel to the long axis of the leaf, in lines; prophyll single, adaxial; leaf blade linear, main venation parallel, of two or more size classes, the veins joining successively from the outside at the apex and forming a fimbrial vein, transverse veinlets +, unbranched [leaf blade characters: ?level], vein/veinlet endings not free, margins entire, Vorläuferspitze +, base broad, ensheathing the stem, sheath open, petiole 0; inflorescence terminal, racemose; flowers 3-merous [6-radiate to the pollinator], polysymmetric, pentacyclic; P = T = 3 + 3, all with three traces, median T of outer whorl abaxial, aestivation open, members of whorls alternating, [pseudomonocyclic, each T member forming a sector of any tube]; stamens = and opposite each T member [A/T primordia often associated, and/or A vascularized from T trace], anther and filament more or less sharply distinguished, anthers subbasifixed, wall with two secondary parietal cell layers, inner producing the middle layer [monocot type]; pollen reticulations coarse in the middle, finer at ends of grain, infratectal layer granular; G [3], with congenital intercarpellary fusion, opposite outer tepals [thus median member abaxial], placentation axile; compitum +; ovule with outer integument often largely dermal in origin, parietal tissue 1 cell across; antipodal cells persistent, proliferating; seed small to medium sized [mean = 1.5 mg], testal; embryo long, cylindrical, cotyledon 1, apparently terminal [i.e. bend in embryo axis], with a closed sheath, unifacial [hyperphyllar], both assimilating and haustorial, plumule apparently lateral; primary root unbranched, not very well developed, stem-borne roots numerous [= homorhizic], hypocotyl short, (collar rhizoids +); no dark reversion Pfr → Pr; nuclear genome [2C] (0.7-)1.29(-2.35) pg, duplication producing monocot LOFSEP and FUL3 genes [latter duplication of AP1/FUL gene], PHYE gene lost. (Some synapomorphies - almost whatever the immediate sister taxa to monocots might be - are in bold.)
[ALISMATALES [PETROSAVIALES [[DIOSCOREALES + PANDANALES] [LILIALES [ASPARAGALES + COMMELINIDS]]]]]: ethereal oils 0; (trichoblasts in vertical files, proximal cell smaller); raphides + (druses 0); leaf blade vernation supervolute-curved or variants, (margins with teeth, teeth spiny); endothecium develops directly from undivided outer secondary parietal cells; tectum reticulate with finer sculpture at the ends of the grain, endexine 0; septal nectaries + [intercarpellary fusion postgenital].
[PETROSAVIALES [[DIOSCOREALES + PANDANALES] [LILIALES + ASPARAGALES] COMMELINIDS]]]] / core monocots: cyanogenic glycosides uncommon; starch grains simple, amylophobic; leaf blade developing basipetally; epidermis with bulliform cells [?level]; stomata anomocytic, (cuticular waxes as parallel platelets).
Age. The crown-group age of this clade is ca 136 Ma (Foster et al. 2016a: q.v. for details), ca 131 Ma (Janssen & Bremer 2004), (132-)128(-121) Ma in Merckx et al. (2008a), ca 123.5 Ma in Magallón et al. (2015), (134-)127(-119) Ma in Givnish et al. (2016b), and similar estimates in Hertweck et al. (2015), Eguchi and Tamura (2016) and Tang et al. (2016). Broader spreads in Magallón and Castillo (2009: relaxed and constrained penalized likelihood datings) are ca 150 Ma and 124 Ma, (133-)124, 109(-99) Ma in Bell et al. (2010), (127-)121, 109(-103) Ma in Wikström et al. (2001: c.f. topology), and 133-108 Ma in Mennes et al. (2013, see also 2015). Note that ages suggested by Lutzoni et al. (2018) for this clade (Petrosavidae) are for its origin, i.e., for the next node down.
Evolution: Divergence & Distribution. Branching in this general part of the tree - i.e. the Petrosaviales, [Dioscoreales + Pandanales], and Liliales clades, and also in Petrosaviales itself, may be somewhere around 125-120 Ma (ca 111 Ma in Bremer 2000b), and the stem groups of all other orders, including those in the commelinid group, have diverged by ca 115 Ma or soon afterwards (Janssen & Bremer 2004) or ca 106 Ma (Magallón et al. 2015). Much of this divergence may have taken place in Southern Gondwana, i.e. Antarctica, Australia, and southern South America (Bremer & Janssen 2006).
For a discussion on leaf development in monocots, see the Acorales page.
Phylogeny. Relationships between commelinids, Asparagales, Dioscoreales, Liliales, and Pandanales were unclear for some time. In a smallish early study, Liliales were sometimes embedded in Asparagales (Eguiarte et al. 1994). An early three-gene (rbcL, atpB, 18S RNA) study (Chase et al. 2000a) showed a polytomy of Petrosaviaceae, Dioscoreales, Pandanales, Liliales, Asparagales and commelinids, although a single shortest tree showed a structure with the taxa in the sequence followed here for some years, i.e. [Petrosaviales [[Pandanales + Dioscoreales] [Liliales [Asparagales + commelinids]]]]; another analysis with placeholders for taxa missing some sequences gave a similar structure, except that Pandanales and Liliales were sister taxa. Note that a combined morphological plus molecular tree in the same volume as the article by Chase et al., Stevenson et al. (2000), suggested a substantially different set of relationships, but bootstraps were not given. Fay et al. (2000a) also suggested a sister relationship between Asparagales and commelinids, although sampling outside Asparagales was sketchy since it was outside their immediate interest. Hilu et al. (2003: matK) i.a. suggested that Orchidaceae might be separate from other Asparagales (the latter being sister to commelinids) and that Dioscoreales and Pandanales formed a clade.
However, a two-gene (matK, rbcL) study (Tamura et al. 2004a) had begun to clarify the situation. Petrosaviaceae (both genera - but see below for a possible third - were studied) were sister to a clade [[Dioscoreaceae + Pandanaceae] [Liliales [Asparagales + commelinids]]] (see Chase et al. 2000a above). Support was quite high (³85% bootstrap) for all order and family branches, although rather lower for [Asparagales + commelinids] (68%) (see also Tamura et al. 2004b, a smaller study; Lam et al. 2016). Davis et al. (2004) also found Petrosaviales to be sister to the same monocots, but with moderate to weak (>72%) support. Givnish et al. (2005: ndhF gene alone) found very much the set of relationships in the tree here, although Pandanales grouped with Liliales (low support) and Dasypogonaceae were sister to [Commelinales + Zingiberales]; a grouping [Liliales [Pandanales + Dioscoreales]] also appeared - and had moderate support - in MP, but not in ML analyses of plastid genomes in Barrett et al. (2013: sampling). Graham et al. (2006) in a study analysing considerable amounts of data also recovered relationships similar to those suggested by Tamura et al. (2004a), all sister taxon relationships in this area having 94% or more support, although that for [Liliales [commelinids + Asparagales]] was only 70% (see also Givnish et al. 2006b; Chase et al. 2006; Magallón et al. 2015). Dioscoreales and Pandanales are sister taxa in most studies (e.g. Hilu et al. 2003; Chase et al. 2006; Qiu et al. 2010: support strong; Magallón et al. 2015; Hertweck et al. 2015; Lam et al. 2015).
On the other hand, G. Petersen et al. (2006b) found trees based on analyses of mitochondrial data in general to be rather incongruent with those based on plastid data, for instance, Orchidaceae grouped with Dioscoreaceae and Thismia, and the positions of Liliales, Asparagales and Dasypogonaceae in particular were very labile. Although G. Petersen et al. (2006b) suggested that the incongruences "could equally well refute the phylogenies based on plastid data" (Petersen et al. 2006b: p. 59), this seems unlikely to happen; idiosyncracies in how the mitochondrial genome - and the atp1 gene seems especially problematical - evolves seems a more likely explanation (see Petersen et al. 2015a for detailed discussion and literature: focus on Acoraceae), and anyhow now one has to keep one's eyes on analyses using data from the nuclear genome.
Analyses using complete plastomes sometimes yielded the clade [Liliales [Pandanales + Dioscoreales]], especially when fewer genes were included in the analyses (Liu et al. 2012; see also Ruhfel et al. 2014: chloroplast genomes, only one species of each included), indeed, a variety of relationships were found in the various analyses carried out by Liu et al. (2012), including Alismatales embedded in the commelinids. For other suggestions of relationships in this area, see Fiz-Palacios et al. (2011).
There was perhaps still uncertainty. In some reconstructions Dioscoreales and Pandanales are adjacent along the monocot spine (e.g. Janssen & Bremer 2004; Bremer & Janssen 2006; Givnish et al. 2006b: not strongly supported), or Nartheciaceae link with Pandanales, the combined group in turn joining with Liliales (Davis et al. 2004: summary of earlier literature on relationships of the two). A four-gene mitochondrial tree suggested the relationships [Asparagales [[Dioscoreales + Pandanales] [Liliales + Commelinids]]], but support was not strong (Qiu et al. 2010), while Davis et al. (2013) recovered a very weakly supported topology [Asparagales s. str. [Orchidaceae + Liliaceae]] in parsimony but not in maximum likelihood analyses (see also Barrett & Davis 2011). Ruhfel et al. (2014) found a [Liliales [Dioscoreales + Pandanales]] clade while in analyses of the nuclear PHYC gene alone Orchidaceae were not sister to other Asparagales - relationships were [Asparagales [Liliales [Orchidaceae + commelinids]]], but support was weak (Hertweck et al. 2015). Recent work using nuclear genomes suggests the grouping [[Asparagales + Liliales] commelinids] (W. J. Baker et al. 2021a: see the Seed Plant Tree of Life, i.2022 and v.2023 versions - however, in the ix.2024 version Orchidaceae were separated from the other Asparagales; H.-T. Li et al. 2021; Timilsena et al. 2022; Zuntini et al. 2024).
Although Petrosaviales are commonly placed as in the tree here, ...[Petrosaviales [Dioscoreales + Pandanales]... (e.g. Lam et al. 2018), until v.2019 no association of Petrosaviaceae with any other order had been strongly supported, hence its inclusion below as a monofamilial Petrosaviales. However, H.-T. Li et al. (2019: plastomes) found quite good (90% bootstrap) support for Petrosaviaceae as sister to Orchidaceae (Asparagales), and another recent plastome study recovered a [Petrosaviales + commelinid] clade (Gitzendanner et al. 2018a, b). Movement of the family awaits confirmation from the nuclear compartment, but initial findings suggested that no movement would be necessary (see W. J. Baker et al. 2021, version ix.2024, etc.; Timilsena et al. 2022; also the comprehensive plastome analysis in Li et al. 2021), but in Zuntini et al. (2024) the relationships ...[[Dioscoreales + Pandanales] [Petrosaviales [[Liliales + Asparagales] [commelinids]]]] were recovered, while in Y.-L. Qiu et al. (2024) the corresponding relationships are ...[Petrosaviales [[Pandanales + Dioscoreales] [Liliales [Asparagales + commelinds]]]].
Some holomycoheterotrophic taxa, e.g., Petrosavia below, cause problems, and in an analysis using three chloroplast genes (not present in all the mycoheterotrophs), branch length could be extremely long, although by no means always (Lam et al. 2016), in some analyses holomycoheterotrophs from three orders (Petrosaviales, Dioscoreales, Liliales) grouping together and forming a clade (Lam et al. 2018). Neyland (2002) found that Thismia was sister to a well supported Burmannioideae, but with less support, but Burmanniaceae s.l. did not link with other Dioscoreales. Analysis of 26S rDNA sequences suggested that Corsiaceae were polyphyletic; Arachnitis perhaps being sister to Thismia and/or Burmannia (Neyland & Hennigan 2003; G. Petersen et al. 2006b: combined analysis). A recent analysis of plastid loci also failed to include Arachnitis in Liliales, and perhaps it was to be included in the commelinids (Kim et al. 2012). However, these relationships have not been confirmed, and these mycoheterotrophs seem to be finding stable resting places, as in studies by Lam et al. (2016, 2018) that included a very wide variety of mycoheterotrophs from the next five orders in the sequence here. However, relationships within Dioscoreaceae, which includes a number of mycoheterotrophic taxa, do not inspire total confidence. Baker et al. (2021: see Seed Plant Tree of Life), looking at 347 nuclear genes, found that Burmannia, normally included in Dioscoreales, was sister to [Dioscoreales + Pandanales], and although Burmannia includes several mycoheterotrophic taxa, the two species examined by Baker et al. were both chlorophyllous. The bizarre mycoheterotrophic Thismia and its relatives were not included in this study, nor were Taccaceae (= Tacca alone). However, in the January 2022 and September 2024 versions, although Thismia was still not included, sampling otherwise was improved and the circumscription of Pandanales was back to normal. Zuntini et al. (2024) included only Thisma rodwayi from the Thisma/Afrothismia clade, and it was sister to the whole [Dioscoreales + Pandanales] clade, albeit with poor support; Burmanniaceae were included in Dioscoreales (q.v.).
For further discussion of relationships in the monocots, see especially the Poales and Acorales pages.
PETROSAVIALES Takhtajan - Main Tree.
Just the one family, 2 genera, 3 species.
Note: In all node characterizations, boldface denotes a possible apomorphy, (....) denotes a feature the exact status of which in the clade is uncertain, [....] includes explanatory material; other text lists features found pretty much throughout the clade. However, the precise node to which many characters, particularly the more cryptic ones, should be assigned is unclear. This is partly because homoplasy is very common, in addition, basic information for all too many characters is very incomplete, frequently coming from taxa well embedded in the clade of interest and so making the position of any putative apomorphy uncertain. Then there are the not-so-trivial issues of how character states are delimited and ancestral states are reconstructed (see above).
Evolution: Divergence & Distribution. This is not exactly a clade in which there has been much diversification (Hertweck et al. 2015).
Includes Petrosaviaceae.
Synonymy: Miyoshiales Nakai - Petrosaviineae Shipunov
PETROSAVIACEAE Hutchinson
Stem with a ring of bundles; sieve tube plastids also with polygonal protein crystalloids; plant glabrous; leaves spiral; inflorescence tacemose, bracts +, bracteoles sublateral (0); T members with a single trace, whorls slightly differentiated, outer somewhat smaller, tube at most short; A inserted at base of T or free, basifixed; microsporogenesis simultaneous; septal nectaries +; G partly connate, opposite outer T, plicate, fusion (congenital and) postgenital, styluli +, short; megaspore tetrads T-shaped; T persistent in fruit; seed endotestal; embryo small/minute; x = 7 (?6, ?14), nuclear genome [1 C] (0179-)1.879(-19.748) pg; seedling?
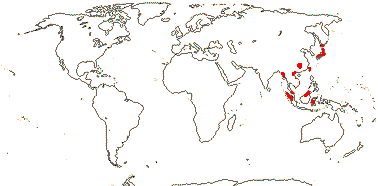
2[list]/3. Scattered, E. India and East Asia to W. Malesia. Map: from Jessop (1979) and Remizowa et al. (2017).
Age. Crown group Petrosaviaceae date to ca 123 Ma (Janssen & Bremer 2004) or (134-)99, 89(-62) Ma (Bell et al. 2010); estimates were (102-)108(-87) Ma in Merckx et al. (2008a), 96-8 Ma and 91-9 Ma in Mennes et al. (2013, 2015 respectively), (115-)108(-99) or (64-)60(-57) Ma in Hertweck et al. (2015), (118.8-)59.1(-17) Ma in Eguchi and Tamura (2016: see discussion) and (119.5-)67(-23) Ma in Givnish et al. 2016b).
1. Japonolirion osense Makino & Tatewaki —— Synonymy: Japonoliriaceae Takhtajan
Leaves on rhizome scaly; pollen gemmate; G stipitate, with postgenital fusion only in centre, ?compitum, stylulus solid, stigma decurrent; ovules 4-12/carpel; fruit septicidal; endotegmen crushed, contents persist; first division of endosperm nucleus symmetrical; n = 15.
1/1. Japan.
2. Petrosavia Beccari —— Synonymy: Miyoshiaceae Nakai
Plant echlorophyllous, holomycoheterotrophic; rhizome with scale leaves; roots axillary, cortex disintegrating [endodermis persists], stele not medullated; stem vascular bundles associated with sclerenchymatous ring; vessels 0; calcium oxalate crystals 0; leaves scales; inflorescence (± corymbose/umbellate); bracteole sublateral; outer T notably smaller than inner; G semi-inferior, stylulus hollow, stigma subcapitate; ovules many/carpel, ana-campylotropous, integumentary obturator +; fruit follicular, also dehiscing abaxially; seeds obliquely arranged, coat loose; first division of endosperm nucleus asymmetrical; n = 12, 13, chloroplast ndh genes 0.
1/2. Eastern India, Japan, southern China to west Malesia.
Evolution: Divergence & Distribution. Remizowa (2011) suggested that the position of the septal nectaries in both the ascidiate and plicate zone of the gynoecium might be unique and so a synapomorphy for this tiny but quite heterogeneous clade.
Plant-Bacterial/Fungal Associations. Roots of the autotrophic and rather uncommon Japonolirion osense from Japan have about 22 phylotypes of Glomales associated with it (Paris-type mycorrhizae), while there was just a single one of these in the much more widely distributed mycoheterotrophic Petrosavia sakuraii, but sampled only from Japan (Yamato et al. 2014); it would be interesting to see what other fungi might be associated with the latter elsewhere in its extensive range. The mycorrrhizae of Petrosavia are of the Paris type (Imhof et al. 2013). For more on mycoheterotrophy in general, see elsewhere.
Genes & Genomes. For evolution (loss of genes, etc.) in the plastome of the mycoheterotroph Petrosavia, see Logacheva et al. (2014; also Luo et al. 2016; Lin et al. 2017); there has also been extensive rearrangment of the genome there.
Chemistry, Morphology, etc.. The roots of Petrosavia have an unmedullated, four-radiate stele surrounded by a massively thickened endodermis (Tomlinson 1982).
The inflorescence of Petrosavia is sometimes shortly branched at the base. Although the genus is often described as having more or less winged seeds, illustrations in e.g. Bhat et al. (2018) suggest that the testa only very loosely surrounds the embryo rather than the presence of an organized wing.
For general information, see Tamura (1998: Nartheciaceae) and especially Cameron et al. (2003) and Remizowa et al. (2017: seed coat endotestal-endotegmic), for anatomy, see Stant (1970), for pollen, see Caddick et al. (1998), for sieve tube plastids, see Behnke (2003), for floral and inflorescence morphology, see Remizowa et al. (2006a, b) and Tobe (2008), and for Petrosavia, see Merckx et al. (2013a), general, Imhof et al. (2013: roots, mycorrhizae), and Waterman et al. (2013: pollination), and for its embryology, see Tobe and Takahashi (2009: nice comparative table).
Phylogeny. Isidrogalvia schomburgkiana (Tofieldiaceae!) was placed sister to Petrosavia (L.-Y. Chen et al. 2013) in a study focussing on the aquatic Alismatales - a rather remarkable position that needs confirmation (but see Luo et al. 2016: contamination or misidentification).
Previous Relationships. Petrosaviaceae have often been included in other families. Dahlgren et al. (1985) placed them - along with genera here included in Nartheciaceae and Tofieldiaceae - in Melianthaceae, and while Tamura (1998) recognised a Petrosaviaceae, this also included members of Tofieldiaceae and Nartheciaceae. Petrosaviaceae s. str. (i.e. Petrosavia alone) were placed in Triuridales by Cronquist (1981) and in Triurididae by Takhtajan (1997); the latter included a monogeneric Japonoliriaceae in his Melanthiales-Liliidae.